Open Access Research Journal of Life Sciences
Issn 2783-025x (online).
Open Access Research Journal of Life Sciences (OARJLS) is a Peer Reviewed, Open Access, International Journal. It is a Referred, Indexed, Online International Journa l. Open Access Research Journal of Life Sciences (OARJLS) is published as a Quarterly Journal with 4 issues per year. Open Access Research Journal of Life Sciences (OARJLS) offers fast publication of quality Research and Review articles. Open Access Research Journal of Life Sciences (OARJLS) publishes manuscripts (Original research, review articles, Short communication and letter to editor) on original work, either experimental or theoretical) from all aspects of Life Sciences (Biology, Genetics, Biological Anthropology, Botany, Medical Sciences, Veterinary Sciences, Biochemical Genetics, Biometry, Clinical Genetics, Cytogenetics, Genetic Epidemiology, Genetic Testing, Evolution and Population Genetics, Immunogenetics and Molecular Genetics). The journal also covers ethical issues. To know more details about Open Access Research Journal of Life Sciences (OARJLS) Click here...

Aims and Scope
Open Access Research Journal of Life Sciences (OARJLS) aims to establish itself as a platform for exchanging ideas in new emerging trends in Biology and Allied Applied Sciences etc. It aims to serve as a forum for life scientists and health professionals. The journal publishes original papers on current research and practical programmes, short notes, news items, book reviews, reports of meetings and professional announcements. Constructive criticisms and discussions of published papers and letters of relevance and interest to the readership will be published at the discretion of the Managing Editor. The journal is committed to prompt review, and priority publication is given to manuscripts with novel or timely findings, and to manuscripts of unusual interests. Since inception, Open Access Research Journal of Life Sciences (OARJLS) is continuously publishing original and best quality research articles. To view full Aims and Scope of Open Access Research Journal of Life Sciences (OARJLS) Click here...
Call for Paper
Open Access Research Journal of Life Sciences (OARJLS) invites you to submit your research work via our Online Submission System or through Email at [email protected] . Make sure that the submitted manuscript should not have been submitted or published previously anywhere else for publication. It is strictly advised to submit original and plagiarism free articles only for possible consideration, else they will be rejected without any response. All received manuscripts will go through Double Blind Peer Review and final decision shall be based on the high level of quality, originality and additional contribution to the existing knowledge. Special note : Once your article is submitted to Open Access Research Journal of Life Sciences (OARJLS) , you cannot submit / present this paper anywhere else, unless your article is rejected by Open Access Research Journal of Life Sciences (OARJLS) . Accepted submission will not be withdrawn or be presentable in any other journal / conference / magazine or any media without written permission of Open Access Research Journal of Life Sciences (OARJLS) .
Why Publish with OARJLS?
- International Open Access Journal
- Peer Reviewed Journal with ISSN (under process),
- Fast and Easy Publication of high quality Research and Review Papers
- Certificate of publication to each author at no cost
- Submission via e-mail /online
- Online tracking of articles submitted.
- Open Access to all Articles: Anytime & Anywhere in the world
- Improved visibility of articles to get more citations
- Immediate response to author queries.
- Highly experiences Editorial Board and Reviewer members
Open Access Research Journal of Life Sciences (OARJLS) Policies
Open access policy: Open Access Research Journal of Life Sciences (OARJLS) is an open access journal which means that all content is freely available without charge to the user or his/her institution. Users are allowed to read, download, copy, distribute, print, search, or link to the full texts of the articles in this journal without asking prior permission from the publisher or the author. This is in accordance with the Budapest Open Access Initiative (BOAI) definition of open access. To know more about Open access Click here
Peer-review policy: The manuscript submitted to Open Access Research Journal of Life Sciences (OARJLS) will be reviewed by two suitable experts in respective subject area. The reports of both the reviewers will be considered when deciding on acceptance/revision or rejection of a manuscript. Editor-In-Chief will make the final decision, based on reviewer’s comments. To know more about Peer-review process Click here
Anti-Plagiarism policy: Open Access Research Journal of Life Sciences (OARJLS) has very strict policy against plagiarism. The authors should ensure that they have written entirely original works, and if the authors have used the work and/or words of others that this has been appropriately cited or quoted. To know more about Plagiarism Click here
Copyright policy: All the articles published in Open Access Research Journal of Life Sciences (OARJLS) are distributed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International license . The journal allows the author(s) to hold the copyright of their work (all usages allowed except for commercial purpose).
Author self-archiving policy: Open Access Research Journal of Life Sciences (OARJLS) allows the authors to self-archive pre-print, post-print and publisher’s version of the article in any Open Access Initiative (OAI) compliant repository.

- MyConference
- Institutions

- Life Sciences
Open Access Research Journal of Life Sciences (OARJLS)
Open Access Research Journal Publication
Philippines

Google Scholar

ScienceGate

ISSN Portal

Peer reviewed only

Open access journal
Journal descriptions.
Open Access Research Journal of Life Sciences (OARJLS) is a Peer Reviewed, Open Access, International Journal. It is a Referred, Indexed, Online International Journal. Open Access Research Journal of Life Sciences (OARJLS) is published as a Quarterly Journal with 4 issues per year. Open Access Research Journal of Life Sciences (OARJLS) offers fast publication of quality Research and Review articles. Open Access Research Journal of Life Sciences (OARJLS) publishes manuscripts (Original research, review articles, Short communication and letter to editor) on original work, either experimental or theoretical) from all aspects of Life Sciences (Biology, Genetics, Biological Anthropology, Botany, Medical Sciences, Veterinary Sciences, Biochemical Genetics, Biometry, Clinical Genetics, Cytogenetics, Genetic Epidemiology, Genetic Testing, Evolution and Population Genetics, Immunogenetics and Molecular Genetics). The journal also covers ethical issues.
- Technical Support
- Find My Rep
You are here
Sage launches new open access journal of international life sciences research.
London , UK . SAGE , one of the world’s leading independent and academic publishers, has today announced that it has launched the Journal of International Life Sciences Research ( JILSR ) , a new open access (OA) journal dedicated to the rapid publication of high-quality articles in the form of life sciences research and review papers.
As a companion to the Journal of International Medical Research , which has been in publication for over 40 years, JILSR will similarly help authors from developing countries to get published, but with a focus on life sciences. JILSR will publish articles from a range of disciplines, and the journal is now open for submissions.
Editor-in-Chief, Malcolm Lader of King’s College London, commented that:
“The Journal of International Life Sciences Research builds on the success of its sister journal to facilitate rapid publication of high-quality life sciences research. It provides the same rapid and thorough peer review, followed by in-depth, high-quality technical editing. I look forward to working with SAGE to provide a valuable forum for this research.”
Karen Philips, Editorial Director, SAGE, further remarked on the launch:
“The launch of the Journal of International Life Sciences Research will provide a platform on which to publish life sciences research. At SAGE, one of our core publishing missions is to support the development of important research fields, ensuring the greatest access and dissemination of key research. We look forward to working with the editorial board and seeing the journal develop and grow internationally.”
More information about the journal can be found here .
SAGE Founded 50 years ago by Sara Miller McCune to support the dissemination of usable knowledge and educate a global community, SAGE publishes more than 850 journals and over 800 new books each year, spanning a wide range of subject areas. A growing selection of library products includes archives, data and video. SAGE remains majority owned by our founder and after her lifetime will become owned by a charitable trust that secures the company’s continued independence. Principal offices are located in Los Angeles, London, New Delhi, Singapore and Washington DC.
Journal of International Life Sciences Research is a journal for the rapid publication of high-quality life sciences research and review articles, on a page charge basis. It is a companion publication to the Journal of International Medical Research , which has been publishing for over 40 years.
Contact (media inquiries only)
- Louise Coady, Head of Corporate Communications and Public Affairs: [email protected] / Tel: (+44) (0)7810 807291
- Camille Gamboa, Director of Corporate Communications and Public Affairs: [email protected] / Tel: (+1) 805-410-7441
Harvard Library Is Launching Harvard Open Journals Program
Harvard Library is launching a new initiative called the Harvard Open Journals Program (HOJP), which will help researchers advance scholarly publishing that is open access, sustainable, and equitable. HOJP will provide publishing services, resources, and seed funding to participating Harvard researchers for new academic journals. All journal articles will be entirely free for authors and readers, with no barriers to publish or to access.
Martha Whitehead, Vice President for the Harvard Library and University Librarian, sees the initiative as an important step in championing open access. Whitehead said, “We want to model the original ethos of open access by reducing barriers and enabling the free flow of ideas and knowledge across the research ecosystem and beyond to the public at large.”
The Harvard Open Journals Program will offer publishing and hosting services to help the Harvard community launch new open access journals, or to convert existing journals to open access. The program will offer two support models: an overlay model which takes advantage of open access repositories, such as Harvard’s DASH , and a brand-new academic press model.
Yuan Li, University Scholarly Communication Officer and Director of Open Scholarship and Research Data Services at Harvard Library, pointed out the innovative nature of the program, “It is new for an institution to support faculty in seeking out an academic press to publish a no-fee open access journal and to provide assistance in securing its long-term funding. And offering a repository overlay journal model provides an alternative that appeals to some editorial boards and is gaining traction through initiatives such as Episciences. As we implement and refine this program on our campus, we hope it will inspire other universities to adopt such approaches to supporting barrier-free scholarly publishing.”
The program is a direct response to faculty interest in alternatives to the article-processing-charge model, in which journals charge author-side fees to publish papers open access. It also supports federal requirements that publications resulting from publicly-funded research be open access.
The open access movement in scholarly publishing seeks to grant free and public online access to publications and data. In recent decades, many researchers have become increasingly concerned that commercial rather than scholarly interests are driving the publishing ecosystem. With some publishers charging article processing fees of over $10,000 per article, skyrocketing costs inhibit many researchers and institutions from publishing in these journals. At the same time, research institutions continue to pay high subscription costs, even as their faculty provide editorial and peer review services mainly for free to the publishers. These practices have led to widespread outcry in the scholarly community, and tensions between publishers and editorial boards have led to the latter’s mass resignations .
Scott Edwards, Professor of Organismic and Evolutionary Biology, and a member of the Harvard Library Faculty Advisory Council, applauds the library’s exploration of new models for supporting open access publishing. Edwards said, “In this increasingly challenging publishing ecosystem, the Harvard Open Journals Program is a welcome new approach.”
“These are sustainable and equitable open access publishing models that allow scholars to take control of scholarly communication,” added Li. “I hope that many research-heavy institutions adopt our approach. The first Harvard Open Access policy launched in 2008 has been adopted nationally and internationally, and it would be great to see similar reach.”
Under Harvard’s Open Access policies, Harvard faculty and researchers give the University a nonexclusive, irrevocable right to distribute their scholarly articles for any non-commercial purpose. Stored and preserved in DASH , Harvard Library’s open access repository, these articles are made available to the scholarly community and the public—anyone with an internet connection can read them for free.
Harvard Library is working closely with the Office of the Vice Provost for Research on launching the HOJP program. John Shaw, Vice Provost for Research and Harry C. Dudley Professor of Structural and Economic Geology, is eager to promote the initiative in the suite of programs that support faculty research. Shaw said, “The launch of HOJP provides very encouraging options for removing barriers to making research results open and expanding their reach.”
The Harvard Open Journals Program will be open to all journals with a current Harvard affiliate on the editorial team or editorial board. Student-run journals are also eligible, as long as they are sponsored by a Harvard faculty member or administrator.
In preparing to launch HOJP this summer, Harvard Library is currently seeking input on program details from interested faculty. HOJP will begin accepting applications in the fall from journals and editorial boards. Colleen Cressman, Librarian for Open Publishing, will manage the program and can be reached by email for more information.
- UB Official
Research Journal of Life Science
Announcements, acreditation certificate.

- Mendeley User Guide
- Insert Citation using Mendeley

Journal Index

- For Readers
- For Authors
- For Librarians
Research Journal of Life Science is an open access publishes papers three times a year on April, August and December. The main objective of Research Journal of Life Science is to provide a platform for the international scholars, academicians and researchers to share the contemporary thoughts and innovation in the fields of life science. Research Journal of Life Science aims to promote studies in life science and thus become the leading international journal in life science in the world.
Research Journal of Life Science has been accredited for five years as scientific journal based on Ministry of Education, Culture, Research and Technology of the Republic of Indonesia (SK No. 164/E/KPT/2021, 27 Desember 2021)
Research Journal of Life Science developed for being International Journal and provide postgraduate as a requirement graduation (University of Brawijaya Rector Decision Letter No. 113/UN10/AK/2017).
We accept submission from all over the world. All submitted articles shall never been published elsewhere, original and not under consideration for other publication.
E-ISSN 2355-9926
Vol 10, No 1 (2023): IN PRESS
Table of contents.
RESEARCH JOURNAL of LIFE SCIENCE
Jurnal Universitas Brawijaya - © 2016
Powered by Open Journal System 2.4.7.1


- The Institut in numbers
- Our governance
- Our commitments
- Conference Center
- Innovation and technology transfer
- Strategic plan for 2019-2023
- How to support us
- Why support us?
- We need you
- Disease sheets
- Find in journal
- Press documents
- Resources for medias
- Our Sars-Cov-2 research projects
- All our COVID-19 news
- Our Covid-19 disease fact sheet
- Our response to fake news
- The Institut Pasteur
- Our missions
- Medical Center
- The research journal
- All SARS-CoV-2 / COVID-19 from the Institut Pasteur
- Education center
- Programs and courses
- Startup Awareness
- Housing in Paris
- Cooperation
- International programs
- International calls
- Fellowships and mobility
- Pasteur Network
- Biological Resource Center (CRBIP)
- WOAH Collaborating Centers
- Industry Partnerships
- Investor Partnerships
- The Carnot Label
- Our job offers
- When you arrive
- Why join us?
- Pasteurians and Alumni Network
- Picture Library
- Scientific publications
- Follow the institut Pasteur on Facebook
- Follow the institut Pasteur on LinkedIn
- Follow the institut Pasteur on Twitter
- Follow the institut Pasteur on Youtube
- International
- Public Health

The Institut Pasteur adopts a Charter for Open Access to Scientific Publications
In 2021, the Institut Pasteur produced a Charter for Open Access to Publications. This charter is part of a broader commitment to open science, as reaffirmed in the 2019-2023 Strategic Plan.
In its 2019-2023 Strategic Plan , the Institut Pasteur confirms its long-term strategy to "promote open access to publications and research data." In 2021, in line with this commitment, it published a Charter for Open Access to Scientific Publications. In the charter, the Institut Pasteur calls for all its scientific publications (research articles, reviews, letters, books, conference papers and preprints) to be published as open access from 2021 onwards.
Find out more
Participation in open science is also one of the criteria taken into consideration when assessing scientists and research engineers. All publications for the period under assessment must be submitted to HAL-Pasteur , the open archive for Institut Pasteur publications.
Many research funding bodies, including the European Commission and the French National Research Agency (ANR), require any publications derived from projects they have funded to be made available as open access. To ensure compliance with these open access policies, the Institut Pasteur's scientists are asked to:
- Submit their publications to the HAL-Pasteur open archive.
- Publish with a CC-BY Creative Commons license and no embargo, in other words make articles available free of charge as soon as they are published , so that they can be read and reused by anyone.
For the latter requirement, there are two possibilities. Publications can be published in a fully open access journal or by applying the Rights Retention Strategy . This strategy was developed by several research funding bodies that have joined forces in cOAlition S .
Using the algorithm provided by the French Ministry of Higher Education, Research and Innovation to calculate the open access rate of French publications, the Institut Pasteur has published its open science barometer. As of July 2021, 85.1% of the Institut Pasteur's publications published in 2020 were available as open access.
The Institut Pasteur's commitment to open science began in 2004, the same year as the Berlin Declaration on Open Access to Knowledge. This strategy to promote open access to publications is in line with France's National Plan for Open Science and with ambitions at international level. The aim is to democratize access to the results of honest, quality research and to speed up innovation by facilitating the reuse of results.
In 2021, the Institut Pasteur also pledged its commitment to another area of open science by adopting a policy for managing and sharing research data and software codes .
Contenus liés

The Institut Pasteur adopts a new data management and sharing policy

Strategic Plan: research advances in 2022
Innovative genomic data analysis method leads to the discovery of novel coronavirus families
Subscribe to our newsletter.
- For Readers
- For Authors
- Publications & Resources
Partners in Progress: Publishers and Librarians Support Open Access Publishing
- There are currently no refbacks.
ALA Privacy Policy
© 2024 Core

- STM Journals
- Special Issues
- Conferences
- Editorial Board Members
- Reviewers Board Members
- Advisory Panel
- Indexing Bodies
- For Authors
- For Reviewers
- For Editors
- For Advisory Board
- Special Issue Guidelines
- Peer-Review Policy
- Manuscript Submission Guidelines
- Publication Ethics and Virtue
- Article Processing Charge
- Editorial Policy
- Advertising Policy
- STM Website and Link Policy
- Distribution and dessemination of Research
- Informed consent Policy
"Connect with colleagues and showcase your academic achievements."
"Unleashing the potential of your words"
"Explore a vast collection of books and broaden your horizons."
"Empower yourself with the knowledge and skills needed to succeed."
"Collaborate with like-minded professionals and share your knowledge."
"Learn from experts and engage with a community of learners."
- ICDR Group of Companies

- Training Programs

Research & Reviews : A Journal of Life Sciences
ISSN: 2249-8656
Journal Menu
Editors overview.

Dr. H.Ramasubba Reddy
Institutional Profile Link : http://svb. . . APID Profile View Full Profile
STM Journals, An imprint of Consortium e-Learning Network Pvt. Ltd. A-118, 1st Floor, Sector-63, Noida, U.P. India, Pin – 201301 E-mail: [email protected] (Tel) (+91) 0120- 4781 200 (Mob) (+91) 9810078958, +919667725932
A Comprehensive Review on COVID-19 Evolution and Development
Characterization of Plant Growth Promoting Bacillus subtilis (VBKT5) Isolated from Vermicompost
Estimation of Microbes on Phyllopalne Region of Healthy and Infected Leaf of Ipomoea cairica by Leaf Impression Method
About the Journal
Research & Reviews : A Journal of Life Sciences (rrjols) : 2249-8656(e) is a peer-reviewed hybrid open-access journal launched in 2011 focused on the rapid publication of fundamental research papers on all areas of Life Sciences. View Full Focus and Scope…
Journal Particulars
[email protected]
+91 120 478 1220
Vol-14 Issue-01 2024
Freshwater Resources in Delhi: A Decadal Analysis of Land Use Changes Saloni Sachdeva, Indira P Sarethy Keywords: LULC, E.coli, Water quality, Groundwater, Delhi, GIS

For Subscriber Access
Special Issue
WEBSITE DISCLAIMER
Last updated: 2022-06-15
The information provided by STM Journals (“Company”, “we”, “our”, “us”) on https://journals.stmjournals.com / (the “Site”) is for general informational purposes only. All information on the Site is provided in good faith, however, we make no representation or warranty of any kind, express or implied, regarding the accuracy, adequacy, validity, reliability, availability, or completeness of any information on the Site.
UNDER NO CIRCUMSTANCE SHALL WE HAVE ANY LIABILITY TO YOU FOR ANY LOSS OR DAMAGE OF ANY KIND INCURRED AS A RESULT OF THE USE OF THE SITE OR RELIANCE ON ANY INFORMATION PROVIDED ON THE SITE. YOUR USE OF THE SITE AND YOUR RELIANCE ON ANY INFORMATION ON THE SITE IS SOLELY AT YOUR OWN RISK.
EXTERNAL LINKS DISCLAIMER
The Site may contain (or you may be sent through the Site) links to other websites or content belonging to or originating from third parties or links to websites and features. Such external links are not investigated, monitored, or checked for accuracy, adequacy, validity, reliability, availability, or completeness by us.
WE DO NOT WARRANT, ENDORSE, GUARANTEE, OR ASSUME RESPONSIBILITY FOR THE ACCURACY OR RELIABILITY OF ANY INFORMATION OFFERED BY THIRD-PARTY WEBSITES LINKED THROUGH THE SITE OR ANY WEBSITE OR FEATURE LINKED IN ANY BANNER OR OTHER ADVERTISING. WE WILL NOT BE A PARTY TO OR IN ANY WAY BE RESPONSIBLE FOR MONITORING ANY TRANSACTION BETWEEN YOU AND THIRD-PARTY PROVIDERS OF PRODUCTS OR SERVICES.
PROFESSIONAL DISCLAIMER
The Site can not and does not contain medical advice. The information is provided for general informational and educational purposes only and is not a substitute for professional medical advice. Accordingly, before taking any actions based on such information, we encourage you to consult with the appropriate professionals. We do not provide any kind of medical advice.
Content published on https://journals.stmjournals.com / is intended to be used and must be used for informational purposes only. It is very important to do your analysis before making any decision based on your circumstances. You should take independent medical advice from a professional or independently research and verify any information that you find on our Website and wish to rely upon.
THE USE OR RELIANCE OF ANY INFORMATION CONTAINED ON THIS SITE IS SOLELY AT YOUR OWN RISK.
AFFILIATES DISCLAIMER
The Site may contain links to affiliate websites, and we may receive an affiliate commission for any purchases or actions made by you on the affiliate websites using such links.
TESTIMONIALS DISCLAIMER
The Site may contain testimonials by users of our products and/or services. These testimonials reflect the real-life experiences and opinions of such users. However, the experiences are personal to those particular users, and may not necessarily be representative of all users of our products and/or services. We do not claim, and you should not assume that all users will have the same experiences.
YOUR RESULTS MAY VARY.
The testimonials on the Site are submitted in various forms such as text, audio, and/or video, and are reviewed by us before being posted. They appear on the Site verbatim as given by the users, except for the correction of grammar or typing errors. Some testimonials may have been shortened for the sake of brevity, where the full testimonial contained extraneous information not relevant to the general public.
The views and opinions contained in the testimonials belong solely to the individual user and do not reflect our views and opinions.
ERRORS AND OMISSIONS DISCLAIMER
While we have made every attempt to ensure that the information contained in this site has been obtained from reliable sources, STM Journals is not responsible for any errors or omissions or the results obtained from the use of this information. All information on this site is provided “as is”, with no guarantee of completeness, accuracy, timeliness, or of the results obtained from the use of this information, and without warranty of any kind, express or implied, including, but not limited to warranties of performance, merchantability, and fitness for a particular purpose.
In no event will STM Journals, its related partnerships or corporations, or the partners, agents, or employees thereof be liable to you or anyone else for any decision made or action taken in reliance on the information in this Site or for any consequential, special or similar damages, even if advised of the possibility of such damages.
GUEST CONTRIBUTORS DISCLAIMER
This Site may include content from guest contributors and any views or opinions expressed in such posts are personal and do not represent those of STM Journals or any of its staff or affiliates unless explicitly stated.
LOGOS AND TRADEMARKS DISCLAIMER
All logos and trademarks of third parties referenced on https://journals.stmjournals.com / are the trademarks and logos of their respective owners. Any inclusion of such trademarks or logos does not imply or constitute any approval, endorsement, or sponsorship of STM Journals by such owners.
Should you have any feedback, comments, requests for technical support, or other inquiries, please contact us by email: [email protected] .
- Reviewer Login |

Journal of Innovative Research in Life Sciences (JIRLS) is an open access, peer reviewed online and print journal, aimed at publishing latest research findings in life sciences. The Journal comprise techniques suitable in promoting the dissemination of research findings that is expected to be of benefit to the basic needs in health, agriculture, biotechnology, pharmaceutical and food industries. JIRLS publishes life science articles of evidence based practices, thus improving the quality of life.
JIRLS is an inherently life sciences research journal as such is constrained to life science research methods only. The journal, JIRLS cut across subject areas like Biochemistry, Biology, Microbiology, Botany, Zoology, Biostatistics, Veterinary science, Health sciences, Agricultural sciences, Bioinformatics, Biotechnology and related fields.
2024, Vol: 6, Issue: 1
- STUDIES ON PREVALENCE OF SHIGELLA AND SALMONELLA SPECIES ON THE FOMITES OF SPECIALIST HOSPITAL, SOKOTO, NIGERIA Keta J.N., Bello I.M., Mubarak A., Keta N.M. JIRLS. 2024; 6(1): 1-11 » Abstract » PDF

Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- My Account Login
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 24 April 2024
Phylogenomics and the rise of the angiosperms
- Alexandre R. Zuntini ORCID: orcid.org/0000-0003-0705-8902 1 na1 ,
- Tom Carruthers 1 na1 ,
- Olivier Maurin ORCID: orcid.org/0000-0002-4151-6164 1 ,
- Paul C. Bailey ORCID: orcid.org/0000-0002-4650-9668 1 ,
- Kevin Leempoel ORCID: orcid.org/0000-0001-7335-7930 1 ,
- Grace E. Brewer 1 ,
- Niroshini Epitawalage 1 ,
- Elaine Françoso ORCID: orcid.org/0000-0002-6464-1240 1 , 2 ,
- Berta Gallego-Paramo ORCID: orcid.org/0000-0002-2016-7161 1 ,
- Catherine McGinnie 1 ,
- Raquel Negrão ORCID: orcid.org/0000-0002-4758-8038 1 ,
- Shyamali R. Roy 1 ,
- Lalita Simpson 3 ,
- Eduardo Toledo Romero 1 ,
- Vanessa M. A. Barber 1 ,
- Laura Botigué ORCID: orcid.org/0000-0001-7114-5168 4 ,
- James J. Clarkson 1 ,
- Robyn S. Cowan 1 ,
- Steven Dodsworth ORCID: orcid.org/0000-0001-6531-3540 5 ,
- Matthew G. Johnson ORCID: orcid.org/0000-0002-1958-6334 6 ,
- Jan T. Kim 7 ,
- Lisa Pokorny ORCID: orcid.org/0000-0002-2478-8555 1 , 8 ,
- Norman J. Wickett ORCID: orcid.org/0000-0003-0944-1956 9 ,
- Guilherme M. Antar ORCID: orcid.org/0000-0001-8109-4544 10 , 11 ,
- Lucinda DeBolt 12 ,
- Karime Gutierrez 12 ,
- Kasper P. Hendriks ORCID: orcid.org/0000-0003-0245-8368 13 , 14 ,
- Alina Hoewener ORCID: orcid.org/0009-0001-6938-9312 15 ,
- Ai-Qun Hu ORCID: orcid.org/0000-0001-9564-878X 1 ,
- Elizabeth M. Joyce ORCID: orcid.org/0000-0001-8291-8058 3 , 16 ,
- Izai A. B. S. Kikuchi ORCID: orcid.org/0000-0002-0258-1537 17 ,
- Isabel Larridon ORCID: orcid.org/0000-0003-0285-722X 1 ,
- Drew A. Larson ORCID: orcid.org/0000-0002-7557-9999 18 ,
- Elton John de Lírio ORCID: orcid.org/0000-0002-9986-9640 10 ,
- Jing-Xia Liu 19 ,
- Panagiota Malakasi 1 ,
- Natalia A. S. Przelomska ORCID: orcid.org/0000-0001-9207-4565 1 , 5 ,
- Toral Shah 1 ,
- Juan Viruel ORCID: orcid.org/0000-0001-5658-8411 1 ,
- Theodore R. Allnutt ORCID: orcid.org/0000-0002-8258-0058 20 ,
- Gabriel K. Ameka ORCID: orcid.org/0000-0001-6659-9982 21 ,
- Rose L. Andrew ORCID: orcid.org/0000-0003-0099-8336 22 ,
- Marc S. Appelhans 23 ,
- Montserrat Arista ORCID: orcid.org/0000-0003-0914-9525 24 ,
- María Jesús Ariza 25 ,
- Juan Arroyo 24 ,
- Watchara Arthan ORCID: orcid.org/0000-0002-6941-2199 1 ,
- Julien B. Bachelier 26 ,
- C. Donovan Bailey 27 ,
- Helen F. Barnes 20 ,
- Matthew D. Barrett ORCID: orcid.org/0000-0002-2926-4291 3 ,
- Russell L. Barrett ORCID: orcid.org/0000-0003-0360-8321 28 ,
- Randall J. Bayer ORCID: orcid.org/0000-0002-7827-5886 29 ,
- Michael J. Bayly ORCID: orcid.org/0000-0001-6836-5493 30 ,
- Ed Biffin ORCID: orcid.org/0000-0002-6582-716X 31 ,
- Nicky Biggs 1 ,
- Joanne L. Birch ORCID: orcid.org/0000-0002-8226-6085 30 ,
- Diego Bogarín ORCID: orcid.org/0000-0002-8408-8841 14 , 32 ,
- Renata Borosova ORCID: orcid.org/0000-0002-3691-5005 1 ,
- Alexander M. C. Bowles ORCID: orcid.org/0000-0002-7487-3811 33 ,
- Peter C. Boyce 34 ,
- Gemma L. C. Bramley 1 ,
- Marie Briggs ORCID: orcid.org/0000-0003-2988-0032 1 ,
- Linda Broadhurst ORCID: orcid.org/0000-0002-9853-3328 35 ,
- Gillian K. Brown 36 ,
- Jeremy J. Bruhl 22 ,
- Anne Bruneau ORCID: orcid.org/0000-0001-5547-0796 37 ,
- Sven Buerki ORCID: orcid.org/0000-0002-8299-6539 38 ,
- Edie Burns 1 ,
- Margaret Byrne ORCID: orcid.org/0000-0002-7197-5409 39 ,
- Stuart Cable 1 ,
- Ainsley Calladine ORCID: orcid.org/0000-0003-4724-8110 31 ,
- Martin W. Callmander ORCID: orcid.org/0000-0003-3641-112X 40 ,
- Ángela Cano ORCID: orcid.org/0000-0002-5090-7730 41 ,
- David J. Cantrill ORCID: orcid.org/0000-0002-1185-4015 20 ,
- Warren M. Cardinal-McTeague ORCID: orcid.org/0000-0003-3558-9794 42 ,
- Mónica M. Carlsen ORCID: orcid.org/0000-0002-1663-0475 43 ,
- Abigail J. A. Carruthers ORCID: orcid.org/0000-0003-2586-9733 1 ,
- Alejandra de Castro Mateo 24 ,
- Mark W. Chase ORCID: orcid.org/0000-0002-9927-4938 1 , 44 ,
- Lars W. Chatrou ORCID: orcid.org/0000-0003-0131-0302 45 ,
- Martin Cheek 1 ,
- Shilin Chen ORCID: orcid.org/0000-0002-0449-236X 46 , 47 ,
- Maarten J. M. Christenhusz ORCID: orcid.org/0000-0003-1398-8743 1 , 48 , 49 ,
- Pascal-Antoine Christin ORCID: orcid.org/0000-0001-6292-8734 50 ,
- Mark A. Clements 35 ,
- Skye C. Coffey ORCID: orcid.org/0000-0001-8211-5546 51 ,
- John G. Conran ORCID: orcid.org/0000-0003-2268-2703 52 ,
- Xavier Cornejo 53 ,
- Thomas L. P. Couvreur ORCID: orcid.org/0000-0002-8509-6587 54 ,
- Ian D. Cowie 55 ,
- Laszlo Csiba 1 ,
- Iain Darbyshire ORCID: orcid.org/0000-0002-5514-9561 1 ,
- Gerrit Davidse 43 ,
- Nina M. J. Davies ORCID: orcid.org/0009-0007-0395-6250 1 ,
- Aaron P. Davis ORCID: orcid.org/0000-0001-9213-4353 1 ,
- Kor-jent van Dijk ORCID: orcid.org/0000-0002-6521-2843 56 ,
- Stephen R. Downie ORCID: orcid.org/0000-0002-4772-0625 57 ,
- Marco F. Duretto ORCID: orcid.org/0000-0003-1013-4291 28 ,
- Melvin R. Duvall 58 ,
- Sara L. Edwards 1 ,
- Urs Eggli ORCID: orcid.org/0000-0002-8842-8188 59 ,
- Roy H. J. Erkens ORCID: orcid.org/0000-0002-1093-0370 14 , 60 , 61 ,
- Marcial Escudero ORCID: orcid.org/0000-0002-2541-5427 24 ,
- Manuel de la Estrella ORCID: orcid.org/0000-0002-4484-3566 62 ,
- Federico Fabriani ORCID: orcid.org/0000-0002-5844-7484 45 ,
- Michael F. Fay ORCID: orcid.org/0000-0003-3491-9093 1 ,
- Paola de L. Ferreira 63 , 64 ,
- Sarah Z. Ficinski 1 ,
- Rachael M. Fowler ORCID: orcid.org/0000-0002-8953-7036 30 ,
- Sue Frisby 1 ,
- Lin Fu ORCID: orcid.org/0000-0003-4933-221X 65 ,
- Tim Fulcher 1 ,
- Mercè Galbany-Casals ORCID: orcid.org/0000-0002-7267-3330 66 ,
- Elliot M. Gardner ORCID: orcid.org/0000-0003-1133-5167 67 ,
- Dmitry A. German ORCID: orcid.org/0000-0001-7951-1644 68 ,
- Augusto Giaretta 69 ,
- Marc Gibernau ORCID: orcid.org/0000-0003-3866-3099 70 ,
- Lynn J. Gillespie ORCID: orcid.org/0000-0003-3129-434X 71 ,
- Cynthia C. González ORCID: orcid.org/0009-0003-3797-1249 72 ,
- David J. Goyder ORCID: orcid.org/0000-0002-3449-7313 1 ,
- Sean W. Graham ORCID: orcid.org/0000-0001-8209-5231 17 ,
- Aurélie Grall 1 ,
- Laura Green ORCID: orcid.org/0000-0003-0774-9498 1 ,
- Bee F. Gunn ORCID: orcid.org/0000-0002-9591-3085 20 ,
- Diego G. Gutiérrez 73 ,
- Jan Hackel ORCID: orcid.org/0000-0002-9657-5372 1 , 74 ,
- Thomas Haevermans ORCID: orcid.org/0000-0001-8934-4544 75 ,
- Anna Haigh ORCID: orcid.org/0000-0003-3435-3501 1 ,
- Jocelyn C. Hall ORCID: orcid.org/0000-0002-5279-4346 76 ,
- Tony Hall 1 ,
- Melissa J. Harrison 3 ,
- Sebastian A. Hatt 1 ,
- Oriane Hidalgo ORCID: orcid.org/0000-0002-1547-8627 77 ,
- Trevor R. Hodkinson ORCID: orcid.org/0000-0003-1384-7270 78 ,
- Gareth D. Holmes ORCID: orcid.org/0000-0003-1120-8731 20 ,
- Helen C. F. Hopkins ORCID: orcid.org/0000-0003-4984-8224 1 ,
- Christopher J. Jackson 20 ,
- Shelley A. James ORCID: orcid.org/0000-0003-1105-1850 51 ,
- Richard W. Jobson ORCID: orcid.org/0000-0002-1822-9634 28 ,
- Gudrun Kadereit 79 ,
- Imalka M. Kahandawala 1 ,
- Kent Kainulainen ORCID: orcid.org/0000-0003-4271-1778 80 ,
- Masahiro Kato ORCID: orcid.org/0000-0001-7752-2745 81 ,
- Elizabeth A. Kellogg ORCID: orcid.org/0000-0003-1671-7447 82 ,
- Graham J. King ORCID: orcid.org/0000-0002-5975-6051 83 ,
- Beata Klejevskaja 84 ,
- Bente B. Klitgaard 1 ,
- Ronell R. Klopper ORCID: orcid.org/0000-0002-0948-5038 85 , 86 ,
- Sandra Knapp ORCID: orcid.org/0000-0001-7698-3945 87 ,
- Marcus A. Koch ORCID: orcid.org/0000-0002-1693-6829 88 ,
- James H. Leebens-Mack ORCID: orcid.org/0000-0003-4811-2231 89 ,
- Frederic Lens ORCID: orcid.org/0000-0002-5001-0149 14 ,
- Christine J. Leon 1 ,
- Étienne Léveillé-Bourret ORCID: orcid.org/0000-0002-0069-0430 90 ,
- Gwilym P. Lewis ORCID: orcid.org/0000-0003-2599-4577 1 ,
- De-Zhu Li ORCID: orcid.org/0000-0002-4990-724X 19 ,
- Lan Li 91 ,
- Sigrid Liede-Schumann ORCID: orcid.org/0000-0003-2707-0335 92 ,
- Tatyana Livshultz ORCID: orcid.org/0009-0000-7244-5955 93 , 94 ,
- David Lorence 95 ,
- Meng Lu ORCID: orcid.org/0000-0002-2921-1632 1 ,
- Patricia Lu-Irving ORCID: orcid.org/0000-0003-1116-9402 28 ,
- Jaquelini Luber 96 ,
- Eve J. Lucas ORCID: orcid.org/0000-0002-7603-435X 1 ,
- Manuel Luján 1 ,
- Mabel Lum ORCID: orcid.org/0000-0002-4712-0603 97 ,
- Terry D. Macfarlane ORCID: orcid.org/0000-0002-7023-9231 51 ,
- Carlos Magdalena 1 ,
- Vidal F. Mansano 96 ,
- Lizo E. Masters ORCID: orcid.org/0000-0001-8632-0758 1 ,
- Simon J. Mayo 1 ,
- Kristina McColl 28 ,
- Angela J. McDonnell ORCID: orcid.org/0000-0002-2549-9253 98 ,
- Andrew E. McDougall ORCID: orcid.org/0000-0001-7713-2800 56 ,
- Todd G. B. McLay ORCID: orcid.org/0000-0001-6405-8007 20 ,
- Hannah McPherson ORCID: orcid.org/0000-0003-0254-4573 28 ,
- Rosa I. Meneses ORCID: orcid.org/0000-0001-8779-2757 99 ,
- Vincent S. F. T. Merckx ORCID: orcid.org/0000-0002-3959-8623 14 ,
- Fabián A. Michelangeli ORCID: orcid.org/0000-0001-7348-143X 100 ,
- John D. Mitchell 100 ,
- Alexandre K. Monro ORCID: orcid.org/0000-0003-4013-3804 1 ,
- Michael J. Moore ORCID: orcid.org/0000-0003-2222-8332 101 ,
- Taryn L. Mueller ORCID: orcid.org/0000-0003-3148-4776 102 ,
- Klaus Mummenhoff ORCID: orcid.org/0000-0002-8449-1593 13 ,
- Jérôme Munzinger ORCID: orcid.org/0000-0001-5300-2702 103 ,
- Priscilla Muriel ORCID: orcid.org/0000-0002-7874-8057 104 ,
- Daniel J. Murphy ORCID: orcid.org/0000-0002-8358-363X 20 ,
- Katharina Nargar ORCID: orcid.org/0000-0002-0459-5991 3 , 35 ,
- Lars Nauheimer ORCID: orcid.org/0000-0002-2847-0966 3 ,
- Francis J. Nge ORCID: orcid.org/0000-0002-0361-8709 31 ,
- Reto Nyffeler ORCID: orcid.org/0000-0002-9293-5967 105 ,
- Andrés Orejuela ORCID: orcid.org/0000-0002-3511-1478 106 , 107 ,
- Edgardo M. Ortiz ORCID: orcid.org/0000-0001-8052-1671 15 ,
- Luis Palazzesi ORCID: orcid.org/0000-0001-8026-4679 73 ,
- Ariane Luna Peixoto 96 ,
- Susan K. Pell ORCID: orcid.org/0000-0002-7214-3225 108 ,
- Jaume Pellicer ORCID: orcid.org/0000-0001-7632-9775 77 ,
- Darin S. Penneys ORCID: orcid.org/0000-0003-0727-2829 109 ,
- Oscar A. Perez-Escobar 1 ,
- Claes Persson 110 ,
- Marc Pignal ORCID: orcid.org/0000-0002-6772-9299 75 ,
- Yohan Pillon ORCID: orcid.org/0000-0003-1760-329X 111 ,
- José R. Pirani ORCID: orcid.org/0000-0001-7984-4457 10 ,
- Gregory M. Plunkett ORCID: orcid.org/0000-0002-0751-4309 100 ,
- Robyn F. Powell 1 ,
- Ghillean T. Prance 1 ,
- Carmen Puglisi ORCID: orcid.org/0000-0003-0304-1812 1 , 43 ,
- Ming Qin 65 ,
- Richard K. Rabeler ORCID: orcid.org/0000-0002-6765-0353 18 ,
- Paul E. J. Rees 1 ,
- Matthew Renner ORCID: orcid.org/0000-0003-2286-7257 28 ,
- Eric H. Roalson ORCID: orcid.org/0000-0003-1655-3681 112 ,
- Michele Rodda ORCID: orcid.org/0000-0002-4130-6685 113 ,
- Zachary S. Rogers ORCID: orcid.org/0000-0003-1136-0960 114 ,
- Saba Rokni 1 ,
- Rolf Rutishauser 105 ,
- Miguel F. de Salas ORCID: orcid.org/0000-0002-7148-639X 115 ,
- Hanno Schaefer ORCID: orcid.org/0000-0001-7231-3987 15 ,
- Rowan J. Schley ORCID: orcid.org/0000-0003-1532-5353 116 ,
- Alexander Schmidt-Lebuhn ORCID: orcid.org/0000-0002-7402-8941 35 ,
- Alison Shapcott ORCID: orcid.org/0000-0003-3734-052X 117 ,
- Ihsan Al-Shehbaz 43 ,
- Kelly A. Shepherd ORCID: orcid.org/0000-0003-1627-7891 51 ,
- Mark P. Simmons 118 ,
- André O. Simões 119 ,
- Ana Rita G. Simões ORCID: orcid.org/0000-0001-7267-8353 1 ,
- Michelle Siros ORCID: orcid.org/0000-0002-4275-1641 1 , 120 ,
- Eric C. Smidt ORCID: orcid.org/0000-0002-1177-1682 121 ,
- James F. Smith 38 ,
- Neil Snow 122 ,
- Douglas E. Soltis ORCID: orcid.org/0000-0001-8638-4137 123 ,
- Pamela S. Soltis ORCID: orcid.org/0000-0001-9310-8659 123 ,
- Robert J. Soreng ORCID: orcid.org/0000-0002-8358-4915 124 ,
- Cynthia A. Sothers ORCID: orcid.org/0000-0002-8052-9590 1 ,
- Julian R. Starr 125 ,
- Peter F. Stevens 43 ,
- Shannon C. K. Straub ORCID: orcid.org/0000-0001-7506-9043 126 ,
- Lena Struwe 127 ,
- Jennifer M. Taylor 91 ,
- Ian R. H. Telford 22 ,
- Andrew H. Thornhill ORCID: orcid.org/0000-0002-0325-5725 22 , 31 , 52 ,
- Ifeanna Tooth 28 ,
- Anna Trias-Blasi 1 ,
- Frank Udovicic ORCID: orcid.org/0000-0003-1697-8444 20 ,
- Timothy M. A. Utteridge ORCID: orcid.org/0000-0003-2823-0337 1 ,
- Jose C. Del Valle ORCID: orcid.org/0000-0001-6023-6208 24 ,
- G. Anthony Verboom 128 ,
- Helen P. Vonow ORCID: orcid.org/0000-0002-3507-4101 31 ,
- Maria S. Vorontsova ORCID: orcid.org/0000-0003-0899-1120 1 ,
- Jurriaan M. de Vos ORCID: orcid.org/0000-0001-6428-7774 129 ,
- Noor Al-Wattar 1 ,
- Michelle Waycott ORCID: orcid.org/0000-0002-0822-0564 31 , 52 ,
- Cassiano A. D. Welker ORCID: orcid.org/0000-0001-6347-341X 130 ,
- Adam J. White 131 ,
- Jan J. Wieringa ORCID: orcid.org/0000-0003-0566-372X 14 ,
- Luis T. Williamson ORCID: orcid.org/0000-0002-0172-6773 56 ,
- Trevor C. Wilson ORCID: orcid.org/0000-0002-9026-0521 28 ,
- Sin Yeng Wong ORCID: orcid.org/0000-0003-4042-9672 132 ,
- Lisa A. Woods 28 ,
- Roseina Woods 1 ,
- Stuart Worboys ORCID: orcid.org/0000-0001-6706-4509 3 ,
- Martin Xanthos 1 ,
- Ya Yang ORCID: orcid.org/0000-0001-6221-0984 133 ,
- Yu-Xiao Zhang 134 ,
- Meng-Yuan Zhou ORCID: orcid.org/0000-0003-1492-8494 19 ,
- Sue Zmarzty 1 ,
- Fernando O. Zuloaga 135 ,
- Alexandre Antonelli ORCID: orcid.org/0000-0003-1842-9297 1 , 110 , 136 , 137 ,
- Sidonie Bellot ORCID: orcid.org/0000-0001-6355-237X 1 ,
- Darren M. Crayn ORCID: orcid.org/0000-0001-6614-4216 3 ,
- Olwen M. Grace ORCID: orcid.org/0000-0003-1431-2761 1 , 106 ,
- Paul J. Kersey ORCID: orcid.org/0000-0002-7054-800X 1 ,
- Ilia J. Leitch ORCID: orcid.org/0000-0002-3837-8186 1 ,
- Hervé Sauquet ORCID: orcid.org/0000-0001-8305-3236 28 ,
- Stephen A. Smith ORCID: orcid.org/0000-0003-2035-9531 18 na2 ,
- Wolf L. Eiserhardt ORCID: orcid.org/0000-0002-8136-5233 1 , 64 na2 ,
- Félix Forest ORCID: orcid.org/0000-0002-2004-433X 1 na2 &
- William J. Baker ORCID: orcid.org/0000-0001-6727-1831 1 , 64 na2
Nature ( 2024 ) Cite this article
35k Accesses
913 Altmetric
Metrics details
- Phylogenetics
- Phylogenomics
- Plant evolution
Angiosperms are the cornerstone of most terrestrial ecosystems and human livelihoods 1 , 2 . A robust understanding of angiosperm evolution is required to explain their rise to ecological dominance. So far, the angiosperm tree of life has been determined primarily by means of analyses of the plastid genome 3 , 4 . Many studies have drawn on this foundational work, such as classification and first insights into angiosperm diversification since their Mesozoic origins 5 , 6 , 7 . However, the limited and biased sampling of both taxa and genomes undermines confidence in the tree and its implications. Here, we build the tree of life for almost 8,000 (about 60%) angiosperm genera using a standardized set of 353 nuclear genes 8 . This 15-fold increase in genus-level sampling relative to comparable nuclear studies 9 provides a critical test of earlier results and brings notable change to key groups, especially in rosids, while substantiating many previously predicted relationships. Scaling this tree to time using 200 fossils, we discovered that early angiosperm evolution was characterized by high gene tree conflict and explosive diversification, giving rise to more than 80% of extant angiosperm orders. Steady diversification ensued through the remaining Mesozoic Era until rates resurged in the Cenozoic Era, concurrent with decreasing global temperatures and tightly linked with gene tree conflict. Taken together, our extensive sampling combined with advanced phylogenomic methods shows the deep history and full complexity in the evolution of a megadiverse clade.
Similar content being viewed by others

Origin of angiosperms and the puzzle of the Jurassic gap
Fossil data support a pre-Cretaceous origin of flowering plants
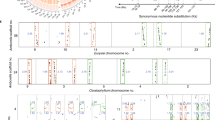

Prickly waterlily and rigid hornwort genomes shed light on early angiosperm evolution
Flowering plants (angiosperms) represent about 90% of all terrestrial plant species 2 but, despite their remarkable diversity and ecological importance underpinning almost all main terrestrial ecosystems, their evolutionary history remains incompletely known. Since their Mesozoic origins 5 , 10 , 11 , angiosperms have had a pervasive influence on the biosphere of Earth, shaping climatic changes at global and local scales 12 , supporting the structure and assembly of biomes 13 and influencing the diversification of other organisms, such as insects, fungi and birds 14 . The evolution of terrestrial biodiversity is thus inextricably linked with the macroevolution of angiosperms, which can only be shown using a robust and comprehensive tree of life. Reconstructing such a tree, however, is challenging because of the sheer diversity of angiosperms and the complex phylogenetic signal in their genomes.
High-throughput DNA sequencing methods now enable us to reconstruct phylogenetic trees that broadly represent the evolutionary signal across entire genomes. Target sequence capture 15 has revolutionized plant phylogenetics by unlocking herbarium specimens as a source of sequenceable DNA 16 , thus removing the chief sampling bottleneck that has obstructed the completion of the tree of life. Although previous work on plants has relied primarily on the widely sequenced plastid genome 3 , 4 , 7 , these technologies now allow us to tap into the evolutionary signal of the much larger and more complex nuclear genome. Universal nuclear probe sets, such as Angiosperms353 (ref. 8 ), have made target sequence capture consistently applicable across broad taxonomic scales, opening doors to collaboration and data integration 17 . As a result, opportunities now present themselves to address fundamental questions in plant evolutionary biology, such as the origin of angiosperms, the tempo and mode of their diversification and the classification of main lineages.
Here, we present a nuclear phylogenomic tree that includes all 64 orders and 416 families of angiosperms recognized by the prevailing classification 18 , using the Angiosperms353 (ref. 8 ) gene panel. Our sampling of 7,923 angiosperm genera (represented by 9,506 species) amounts to a 15-fold increase compared to previous work 9 . Leveraging a dataset of 200 fossil calibrations, we scale the tree to time, effectively capturing evolutionary divergences for all but the most recent 15% of angiosperm history. Although our tree broadly supports relationships predicted by previous studies primarily based on plastid data, it also shows previously unknown relationships and highlights some that remain intractable despite a vast increase in data. Gene tree conflict is tightly linked to diversification across the tree. We find evidence for high levels of conflict associated with an early burst of diversification, which is followed by an extended period of constant diversification rates underpinned by a tapestry of varied lineage-specific patterns. Diversification then increases in the Cenozoic Era, potentially driven by global climatic cooling. Our results highlight the fundamental role of botanical collections in reconstructing the tree of life to illuminate long-standing questions in angiosperm macroevolution.

The angiosperm tree of life
Our phylogenetic tree includes 58% of the approximately 13,600 currently accepted genera of angiosperms (Fig. 1 and Supplementary Table 1 ; ref. 2 ). Together, the 7,923 genera encompass 85.7% of total known angiosperm species diversity. We produced data for 6,777 of these genera; before this study, 3,154 of these lacked publicly available genomic data, of which 393 lacked any form of DNA sequence data. For the remaining genera, data were obtained from public repositories. Sampling for this project was possible thanks to the collaborative effort of many biodiversity institutions from around the world, including 163 herbaria in 48 countries. More than one-third of species were sourced directly from herbarium specimens, some dating back nearly 200 years. Many phylogenetically problematic lineages with unconventional genome evolution were sampled, such as holoparasites, mycoheterotrophs and aquatics. Many of the species included are threatened and four are extinct (or extinct in the wild). The resulting tree of life presented here is one of the largest genomic trees generated yet for angiosperms as a whole.
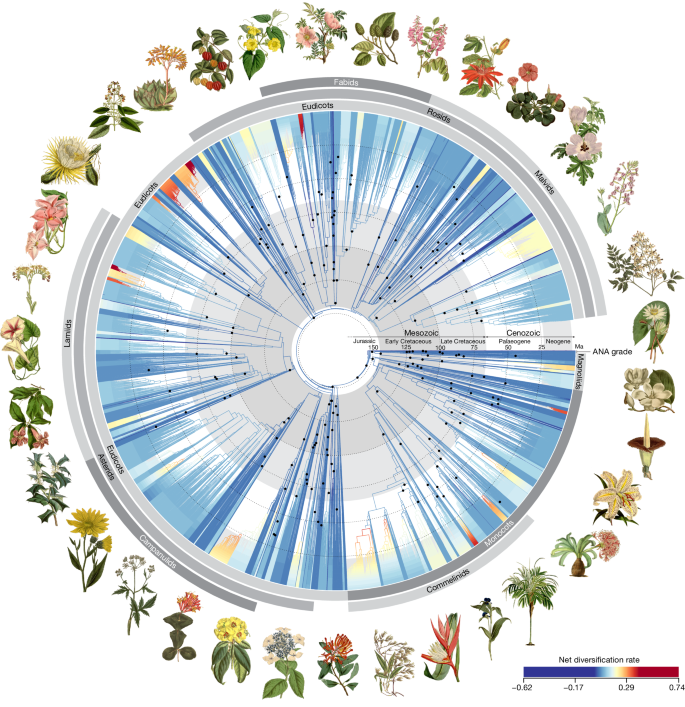
All 64 orders, all 416 families and 58% (7,923) of genera are represented. The young tree is illustrated here (maximum constraint at the root node of 154 Ma), with branch colours representing net diversification rates. Black dots at nodes indicate the phylogenetic placement of fossil calibrations based on the updated AngioCal fossil calibration dataset. Note that calibrated nodes can be older than the age of the corresponding fossils owing to the use of minimum age constraints. Arcs around the tree indicate the main clades of angiosperms as circumscribed in this paper. ANA grade refers to the three consecutively diverging orders Amborellales, Nymphaeales and Austrobaileyales. Plant portraits illustrating key orders were sourced from Curtis’s Botanical Magazine (Biodiversity Heritage Library). These portraits, by S. Edwards, W. H. Fitch, W. J. Hooker, J. McNab and M. Smith, were first published between 1804 and 1916 (for a key to illustrations see Supplementary Table 2 ). A high-resolution version of this figure can be downloaded from https://doi.org/10.5281/zenodo.10778206 (ref. 55 ).
The phylogenomic challenge
Large genomic datasets present challenges to phylogenetic inference. One issue is accurate homology assessment, which proved intractable across the full span of our dataset, even with the most advanced multiple sequence alignment methods. Another challenge is the efficient search of tree space based on gene matrices that have many more taxa than characters. We overcame both challenges with a divide-and-conquer approach (Supplementary Fig. 1 ). First, we computed a backbone species tree with sampling limited to five species per family (1,336 (15%) samples in total) and targeted to represent their deepest nodes (Supplementary Fig. 2 ). We used the backbone species tree to delimit taxon subsets for the construction of order-level gene alignments, which were then merged into global alignments. We then computed global gene trees from the global alignments, using backbone gene trees (inferred during the estimation of the backbone species tree) as topological constraints to reduce tree space while still letting gene trees differ from each other. The smaller number of samples in the backbone dataset permits a more thorough search of tree space, resulting in greater confidence at deeper nodes than could be achieved in an unconstrained global analysis. This approach allows a trade-off between comprehensive sampling and tree search robustness while accommodating putative discordance among gene trees. Finally, we used the global gene trees to generate a global species tree in a multispecies coalescent framework (Supplementary Fig. 3 ).
A widespread concern in phylogenomic analysis is the presence of undetected gene copies. Our findings are unlikely to be affected by this because we used genes that have been selected to be mostly single-copy across green plants 8 , 9 . Although gene duplication cannot be ruled out 19 , the methods we used have been shown to be robust to the presence of paralogues 20 . In addition, a full assessment of orthologues was not computationally tractable but should be undertaken when methods become available to fully unravel the complexity of genome evolution at this scale 21 .
Phylogenetic insights from nuclear data
Our results broadly corroborate the prevailing understanding of angiosperm phylogenetic relationships, which rests on three decades of molecular systematic research largely built on data from the plastid genome 3 , 4 , 18 , 22 . We recover all main lineages of angiosperms, namely Amborellales, Nymphaeales, Austrobaileyales, Ceratophyllales and the three larger clades, monocots, magnoliids (including Chloranthales) and eudicots (Figs. 1 and 2 ). Although some of the relationships among those groups, such as the placement of Amborellales as sister group to all other angiosperms, are well-established and confirmed here, others, such as the placement of Ceratophyllales, which have been unstable in previous work 4 , 9 , remain inconclusive in our results. Despite the contrasting biological properties of the nuclear and plastid genomes (for example, size, copy number, mode of inheritance, recombination and evolutionary rate), which can lead to conflicting phylogenetic results, our findings largely support the mostly plastid-based phylogenetic classification of the Angiosperm Phylogeny Group 18 (Extended Data Fig. 1 ). For example, 58 of the 64 now accepted orders and 406 of the 416 families are recovered as monophyletic (excluding artefacts; Supplementary Table 1 ). The most striking exception is the non-monophyly of Asteraceae, the largest angiosperm family comprising the sunflowers and their relatives. Our tree also confirms 85% of the relationships among families recovered by ref. 4 using plastid genomes (Supplementary Fig. 4 ).
The overall stability of established relationships is unevenly distributed across the tree, as observed in contrasting patterns in the main eudicot clades, the asterids and rosids, which account for 35% and 29% of angiosperm diversity, respectively 2 . The relationships among main orders of asterids are stable 9 , with a clade comprising Ericales and Cornales sister to all other asterids and the remaining 15 orders divided in two main clades (campanulids and lamiids), both long characterized by their contrasting floral ontogeny 23 . Relationships contrasting with the status quo are mostly restricted to small orders, such as the paraphyly of Aquifoliales, Bruniales and Icacinales. These DNA-defined orders were consistently recovered as highly supported clades in plastome analyses 4 , 24 but they lack morphological cohesion. Given their placement in our phylogenetic tree and unique morphologies, these changes, although small, will alter our understanding of the evolution of asterids.
By contrast to asterids, our findings in rosids conflict markedly with plastid-based evidence. First, we resolve Saxifragales, rather than Vitales 4 , as sister to the remainder of rosids. In rosids, the fabid and malvid subclades, recovered as reciprocally monophyletic by plastid data 4 , 22 , are substantially rearranged into a grade of four orders subtending two well-supported sister clades, which we designate here as the recircumscribed fabids and malvids. The new fabid clade (Cucurbitales, Fabales, Fagales and Rosales) has long been characterized by symbiotic nitrogen fixation 25 . In the new malvids (Brassicales, Celastrales, Huerteales, Malpighiales, Malvales, Oxalidales, Picramniales and Sapindales), Oxalidales is resolved as two independent lineages, the core emerging closer to Brassicales, Malvales and Sapindales, whereas Huaceae emerges in the position conventionally occupied by Oxalidales, that is, closer to Malpighiales and Celastrales (the former Celastrales–Oxalidales–Malpighiales (COM) clade 18 ).
Notwithstanding the many well-supported confirmatory and new findings, some key relationships remain contentious and cannot be resolved by our data. These areas of high gene tree conflict often coincide with biological processes that confound phylogenetic inference. For example, the uncertain placements of eudicot orders Caryophyllales, Dilleniales and Gunnerales are probably impacted by key whole genome duplications 9 , 26 . The poor support for relationships among magnoliids, monocots, eudicots and Ceratophyllales might be explained by ancient hybridization events, such as that recently proposed for the origin of the monocots 27 . These examples highlight the importance of areas of poor resolution as waymarkers to biological events meriting further study.
Time frame for angiosperm macroevolution
Our tree was analysed in combination with a dataset of 200 fossil calibrations (originally described in ref. 5 , with modifications) to estimate divergence times and rates of diversification. Because the age of angiosperms is uncertain 28 , we dated the tree with two different maximum constraints at the angiosperm crown node (154 and 247 million years ago (Ma), termed the young tree and old tree, respectively), which reflect realistic upper and lower bounds for the maximum age of this node 5 , 28 . These different constraints affected age estimates across angiosperms (Extended Data Fig. 2 , Supplementary Fig. 5 and Supplementary Table 3 ). For example, in the young tree, stem node age estimates for Nymphaeales, Austrobaileyales and Ceratophyllales were 153, 152 and 152 Ma, respectively, whereas in the old tree the equivalent age estimates were 245, 244 and 243 Ma. Likewise, for larger clades such as magnoliids, monocots and eudicots, crown node age estimates were 151, 149 and 151 Ma in the young tree and 238, 237 and 241 Ma in the old tree. This range in age estimates is consistent with the most comprehensive comparable study 5 (Extended Data Fig. 3 ) but our trees provide age estimates for a further 7,000 nodes. In subsequent analyses, we indicate if differing age estimates between the young tree and old tree cause substantially different interpretations of angiosperm diversification.
With our sampling across angiosperms, we ensured that deeper branching events leading to extant lineages are comprehensively represented, while recognizing that extinct lineages are inaccessible to genomic methods. However, our dated trees are sparsely sampled at the species-level, meaning that branching events are incompletely represented towards the present, limiting diversification inferences in that time window. To address this, we developed a simulation-based approach to quantify the sampling fraction through time. For both dated trees, the lineage representation begins to drop substantially (below 75%) around 50 Ma (Supplementary Fig. 6 ). However, the most dramatic fall in lineage representation occurs in the most recent 20 Myr, in which it falls from around 50% to slightly more than 1% at present. Our investigation of angiosperm diversification should be interpreted with this broader context in mind. In particular, inferences in the most recent 20 Myr may be updated in the future with denser species sampling.
The diversification of angiosperms
Diversification linked to gene conflict.
We used our dated trees to reconstruct both diversification and gene tree conflict across a broad range of temporal and phylogenetic scales and investigate the relationship between them. We show that throughout angiosperm macroevolution, elevated gene tree conflict was tightly associated with elevated diversification. At a general level, this relationship is visible by simply comparing estimated diversification rates with gene tree conflict across all angiosperms through time (Fig. 3a ). Meanwhile, in a branch-specific analysis using the temporal duration of branches as a proxy for the rate at which branches are diversifying, we also show that conflict and diversification rate are positively correlated (Extended Data Fig. 4 ) ( P < 0.001, r 2 = 0.51).
To characterize the theoretical basis of this relationship, we simulated species trees with corresponding gene trees under different diversification scenarios in a multispecies coalescent framework. These simulations showed that gene tree conflict is positively correlated with diversification when caused by incomplete lineage sorting, assuming that effective population size is constant (Supplementary Fig. 7 ). Our empirical results are largely consistent with such a scenario. Other potential causes of gene tree conflict such as whole genome duplication and hybridization may also be associated with rapid diversification and have been recorded extensively throughout angiosperms 29 , 30 . Overall, however, gene tree conflict seems to be reliable corroborating evidence for investigating temporal patterns of angiosperm diversification.
Early burst of angiosperm diversification
Our lineage-through-time (LTT) heatmap and diversification rate estimates through time both indicate an explosive early phase of diversification of extant lineages during the Late Jurassic and Early Cretaceous Periods (Fig. 2b and Fig. 3a ). An early burst of angiosperm diversification, popularized as ‘Darwin’s abominable mystery’ 31 , 32 , is expected given the sudden emergence of diverse angiosperm fossils during the Early Cretaceous 11 , 33 , 34 , 35 . Phylogenetic studies based on single or few genes have also implied that angiosperms diversified rapidly in the Early Cretaceous 7 , 36 , 37 , 38 . Our dated tree corroborates the existence of a distinct early burst of diversification, associated with high levels of gene tree conflict (Fig. 3a and Supplementary Fig. 8 ), further increasing our confidence in this finding.
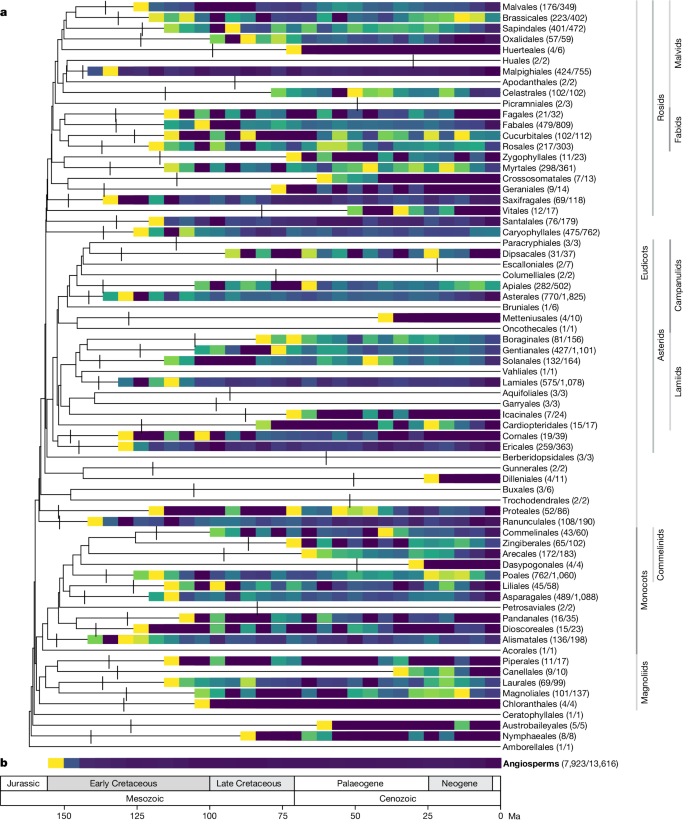
The results illustrated are based on the young tree (maximum constraint at the root node of 154 Ma). a , Time-calibrated summary phylogenetic tree with LTT plots rendered as heatmaps for all orders with four or more sampled genera. The log-transformed increase in the number of lineages is depicted in 5 Myr intervals, omitting crown nodes, which disproportionately altered the visualization; crown node locations are indicated by vertical lines. The yellow to blue colour scale represents steep to shallow slopes. For each order, the numbers of sampled and total genera are provided. b , A global LTT heatmap for all angiosperms is shown at the bottom of the figure as a whole.
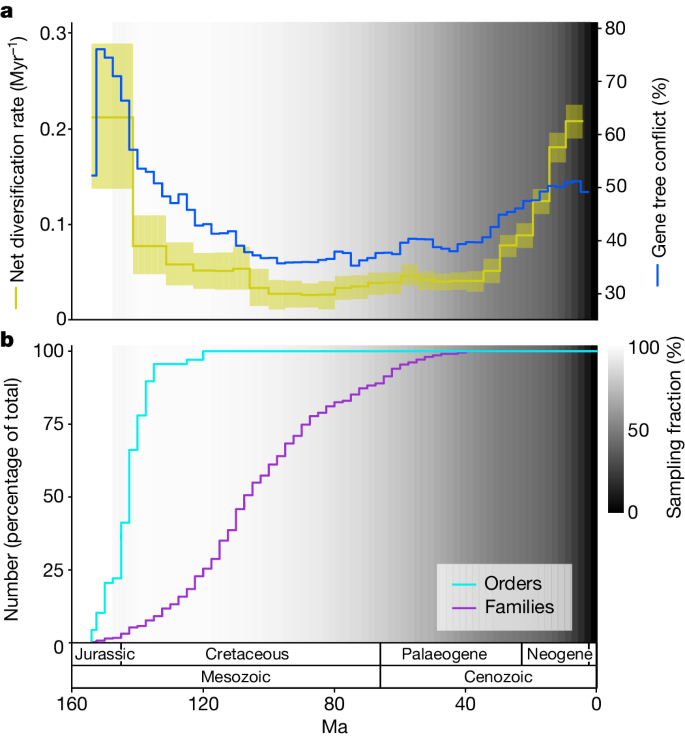
The results illustrated are based on the young tree (maximum constraint at the root node of 154 Ma). See Extended Data Fig. 5 for results based on the old tree. a , Estimated net diversification rate through time (yellow, left y axis) and the level of gene tree conflict through time (blue, right y axis). Net diversification rates are estimated with a model that enables speciation rates to vary between time intervals; the line is the posterior mean and the yellow shaded area is the 95% highest posterior density. Gene tree conflict is calculated from the percentage of gene trees that do not share a congruent bipartition with each species tree branch, with the plotted value being the mean across all species tree branches that cross each 2.5 Myr time slice. b , Cumulative percentage of extant orders and families that have originated through time. In both a and b , the background grey-scale gradient is the estimated percentage of extant lineages represented in the species tree through time (sampling fraction).
More than 80% of extant angiosperm orders originated during the early burst of diversification (Fig. 3b ). Although not strictly comparable because of their subjective delimitation, orders represent the main components of angiosperm feature diversity, which have arisen rapidly after the crown node of angiosperms. In the young tree (Fig. 3 ), the early burst occurs during the Cretaceous, consistent with the hypothesis that a Cretaceous terrestrial revolution was triggered by the establishment of main angiosperm lineages 14 , 39 , 40 . More controversially, the old tree places the early burst in the Triassic Period (Extended Data Fig. 5 ), which is dramatically at variance with the palaeobotanical record 33 , 34 , highlighting that current molecular dating methods are unable to resolve the age of angiosperms 28 .
A tapestry of lineage-specific histories
Following the early burst, overall rates of diversification across angiosperms continued at a lower, constant pace for at least 80 Myr (Fig. 3a ), during which time around three-quarters of all families originated (Fig. 3b ). As expected, this phase of slower diversification was associated with lower levels of gene tree conflict. Despite the constancy of overall rates, diversification during this period was underpinned by a complex tapestry of lineage-specific patterns. This is illustrated by the LTT heatmap, which shows profound differences in diversification trajectories among orders (Fig. 2 ) and by the estimation of around 160 lineage-specific diversification rate shifts in angiosperms, most of which occur during this period. These rate shifts have a widespread phylogenetic distribution, with most orders containing at least one rate shift and many containing several nested shifts (Supplementary Table 4 ). The importance of nested rate shifts is highlighted extensively in discussions of evolutionary radiation 41 , 42 and underpins the continual response of diversification to dynamic extrinsic and intrinsic conditions. However, because these rate shifts are temporally scattered, as also shown by ref. 43 , they do not lead to observable global rate shifts across angiosperms.
A Cenozoic diversification surge
A second surge in angiosperm diversification occurred during the Cenozoic Era (Fig. 3a ). The occurrence of this surge, despite the already high standing diversity of angiosperms at the time, suggests that diversification was unaffected by diversity-dependent processes, that is, the filling of available niche space as clades diversify 44 . Instead, this finding is consistent with previously proposed positive feedbacks between increased diversity and increased rates of diversification in angiosperms 14 , alongside more positive feedbacks, for example, between angiosperm and insect diversification 45 , 46 . Alternatively, global climatic cooling during the Cenozoic acting as a driver of angiosperm diversification could explain this finding 7 , 47 , 48 , 49 . Importantly, an even larger Cenozoic surge would probably be inferred with increased sampling that addresses the under-representation of branching events in the recent time window. The temporal distribution of lineage-specific diversification rate shifts may offer some insight into the cause of the Cenozoic surge. Many of the largest diversification rate increases occur during the Cenozoic, whereas the number of diversification rate decreases declines markedly during this period (Fig. 4 ). These large rate increases may underpin the Cenozoic surge. The expansion of taxon sampling should be given priority to confirm these patterns.
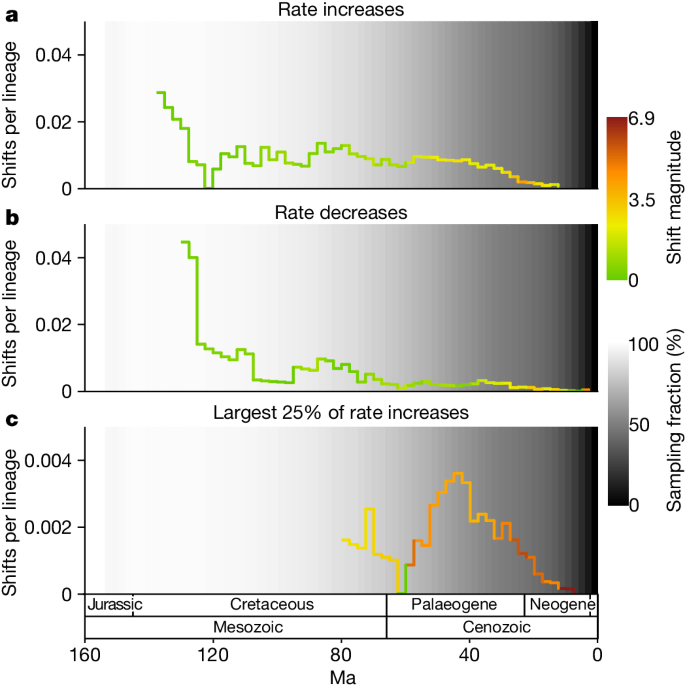
The results illustrated are based on the young tree (maximum constraint at the root node of 154 Ma). See Extended Data Fig. 6 for results based on the old tree. a , Diversification rate increases per LTT. The colour corresponds to the average magnitude of the rate increases during the time period. b , Equivalent to a but for rate decreases. c , Equivalent to a but focusing on the largest 25% of diversification rate increases. In a , b and c , the number of shifts is from the maximum a posteriori shift configuration with the prior for the number of shifts set to 10 and the background grey-scale gradient is the estimated percentage of extant lineages represented in the species tree through time (sampling fraction).
The nuclear phylogenomic framework presented here is the result of an ongoing initiative to complete the tree of life for all angiosperm genera 50 , a milestone in our understanding of angiosperm evolutionary relationships. This study not only sheds light on much of the deep diversification history of the angiosperms but also lays foundations for future work towards a species-level tree 50 . The standardized panel of nuclear genes in our dataset paves the way for more collaborations and data integration 17 , 51 , while the open availability of universal tools to sequence them (that is, Angiosperms353 probes 8 ) has made nuclear genomic data more accessible at relatively low cost. The accelerating uptake of this approach 52 , 53 , 54 , which is readily applicable to herbarium collections 16 , indicates that large volumes of data will soon become available for a wide range of applications in plant diversity, systematic and macroevolutionary research.
Our fossil-calibrated, phylogenomic tree enables a range of unique insights into broad-scale diversification dynamics of angiosperms, substantiating the early burst of diversification anticipated by Darwin while illuminating the complexity and conflict in the lineage histories underlying it. This sets the scene for future research, extending these investigations to shallower phylogenetic scales or digging more deeply into the data to discover the processes driving angiosperm diversification, such as genomic conflict, polyploidy, selection, trait evolution and adaptation. The challenges brought by the scale of this dataset and its ongoing expansion may also catalyse the development of methods which take full advantage of the global proliferation of genomic data.
As part of the Plant and Fungal Trees of Life (PAFTOL) Project at the Royal Botanic Gardens, Kew 50 , we assembled a nuclear genomic dataset consisting of newly generated data and data mined from public repositories. Our objective was to sample at least 50% of all angiosperm genera, with genera selected in a phylogenetically representative manner on the basis of published research. To avoid excessive imbalance in the tree, we included only one sample per species and a maximum of three species per genus. When several samples were available for the same species, we selected those with the largest amount of data, that is, more genes and a higher sum of gene length. For genera with several species available, the criterion for selection was primarily phylogenetic representation followed by amount of data. One species of each gymnosperm family was selected to form the outgroup, totalling 12 samples.
We produced target sequence capture data for 7,561 samples using the universal Angiosperms353 probe set 8 following established laboratory protocols 50 , 56 . We complemented our dataset with publicly available data for 2,054 species, sourced from the One Thousand Plant Transcriptomes Initiative 9 (OneKP; 564 samples), annotated and unannotated genomes (151 samples) and the sequence read archive (SRA; 1,339 samples), the last including transcriptomes (for example, see refs. 57 , 58 ) and target capture data (for example, see refs. 59 , 60 ). To standardize taxonomy and nomenclature, all species names and families were harmonized with the World Checklist of Vascular Plants 2 and orders with APG IV if possible 18 .
Sequence recovery
Sequence recovery was carried out in two ways, depending on the type of input data. For recovery on the basis of raw reads, that is, Angiosperms353 data or data mined from the SRA, we used HybPiper v.1.31 (ref. 61 ), embedded in a bespoke pipeline ( https://github.com/baileyp1/PhylogenomicsPipelines ). Raw reads were trimmed using Trimmomatic 62 to remove low-quality bases and short sequences. In HybPiper, reads were initially binned into genes using BLASTN and an amino acid target file as reference (Supplementary File 1 ). Individual genes were assembled de novo using SPADES 63 and refined by joining and trimming gene contigs to match coding regions using Exonerate 64 . For genes with paralogue warnings, only the putative orthologue as identified by HybPiper was used. Exclusion of genes with several copies per species has been shown to have negligible impact on species tree inference when it is performed under a multispecies coalescent framework, as described below 20 . Conversely, the inclusion of several copies per species would have rendered our study computationally intractable. Gene sequences from assembled genomes and OneKP transcriptomes were recovered using custom scripts described in ref. 50 . Briefly, the assembled sequences were searched against the target file mentioned above using BLASTN, selecting the best match for each gene and trimming it to the BLAST hit. For a few Angiosperms353 samples that represented the sole accession of their respective families ( Ixonanthes reticulata , Mitrastemon matudae and Tetracarpaea tasmannica ) and had poor recovery from HybPiper (that is, below 5 kilobase pairs (kb) in total sum of contig length), recovery was undertaken following ref. 50 , using less stringent recovery thresholds. The average recovery per order is presented in Supplementary Fig. 9 .
Phylogenetic inference
To analyse the dataset, we devised a divide-and-conquer approach. First, we computed a backbone tree, sampling up to five species per family, to test the monophyly of orders and to rigorously explore deep relationships. We used the backbone tree to identify groups (orders or groups of orders) for multiple sequence alignment, with the aim of producing refined subalignments among closely related taxa. Subsequently, the subalignments were merged into global gene alignments and global gene trees were inferred from these using the respective gene trees from the backbone analysis as constraints. Finally, we inferred a multispecies coalescent tree using the estimated gene trees. The inference pipeline is summarized in Supplementary Fig. 1 .
Backbone tree inference
The samples for the backbone were selected so as to represent the crown node and deepest divergences in each family. For families with five or fewer samples (279 families), all samples were included. For those with more than five samples (156), we selected the best sample (most genes and longest sequence) of each consecutively diverging clade (based on published phylogenetic evidence and preliminary analyses of our own data), until five samples were included. To evaluate the extent to which sample selection might affect the backbone tree topology, we inferred 20 backbone replicates, randomly selecting five samples for each family with more than five samples (among the 50% best samples in terms of gene number and gene length recovered). We then summarized the trees to family level and computed Robinson–Foulds distances between the backbone and the 20 replicates (Supplementary Fig. 10 ).
The phylogenetic reconstruction of the backbone involved up to two iterations of gene alignment and gene tree estimation, with an intermediate step of outlier removal. This was followed by species tree inference in a multispecies coalescent framework. In the first iteration, all sequences for a given gene were aligned using MAFFT v.7.480 (ref. 65 ) (with ffnsi method, that is, --retree 2 --maxiterate 1000) and with the direction of the sequence adjusted (--adjustdirection). After removing sites with more than 90% missing data with Phyutility 66 , gene trees were estimated using IQ-TREE v.2.2.0-beta 67 , keeping identical sequences in the analysis (--keep-ident), setting the substitution model to GTR + G and estimating branch support with 1,000 ultrafast bootstrap replicates 68 . Before the second iteration, we identified long branch outliers using TreeShrink 69 in ‘all-genes’ mode and rerooting at the centroids of the trees. A second iteration of gene alignment, removal of gappy sites and gene tree estimation was performed for genes with outliers after the removal of outlier sequences. Subsequently, the resulting gene trees were summarized into a species tree using ASTRAL III v.5.7.3, a quartet-based species tree estimation method statistically consistent with the multispecies coalescent model 70 , enabling the full annotation option (-t 2), having first collapsed poorly supported nodes (ultrafast bootstrap ≤ 30%) in the input gene trees using Newick utilities 71 .
Order-level subalignments
For the order-level subalignments, most orders were analysed individually, following the same method described for the backbone. In some cases, smaller orders (fewer than 50 samples) were analysed together with larger ones if they formed monophyletic groups in the backbone. These groups are: (1) Commelinales with Zingiberales, (2) Dioscoreales with Pandanales, (3) Fagales with Fabales, (4) Columelliales, Dipsacales, Escalloniales and Paracryphiales with Apiales, (5) all magnoliids (Canellales, Laurales, Magnoliales and Piperales) and (6) all gymnosperms together (Cycadales, Ephedrales, Gnetales, Ginkgoales and Pinales). Conversely, orders emerging as non-monophyletic in the backbone were split into monophyletic subgroups as follows: (1) Cardiopteridaceae and Stemonuraceae separate from the rest of Aquifoliales, (2) Dasypogonaceae separate from the rest of Arecales, (3) Collumelliaceae separate from the rest of Bruniales, (4) Oncothecaceae separate from the rest of Icacinales and (5) Huaceae separate from the rest of Oxalidales. The groupings of samples used in the order-level subalignments are provided in Supplementary Table 1 . Very small groups, comprising one or two samples (termed orphan sequences), were not included in subalignments and were incorporated directly in global analyses.
Global gene alignments and trees
We produced global gene alignments by merging the order-level subalignments (before removal of gappy sites) and adding the orphan non-aligned sequences using MAFFT 65 , with up to 100 refinement iterations. This approach yields alignment across the order-level subalignments without disrupting the structure in the subalignments. The final gene alignments were cleaned by removing gappy sites. A summary of the alignments was produced with AMAS 72 (Supplementary Table 5 ) and the average occupancy per gene per order is presented in Supplementary Fig. 11 .
We then estimated gene trees in Fasttree v.2.1.10 (ref. 73 ), setting the model to GTR + G, using pseudocounts to avoid biases from fragmentary sequences and increasing search thoroughness (-spr 4, -mlacc 2 and -slownni). We used the gene trees from the backbone analysis to constrain the topology of each respective global gene tree. To avoid propagating error from the backbone analysis to the global analysis, we removed potentially misleading signal from the backbone gene trees before applying them as constraints. First, branches with bootstrap values below 80% were collapsed to avoid enforcing poorly supported relationships. Second, tips placed far from the rest of their order were algorithmically removed (but retained in global gene alignments). Once global gene trees were estimated, outlier long branches were removed using TreeShrink and the set of pruned gene trees was used to compute the global species tree using ASTRAL-MP v.5.15.5 (ref. 74 ), after collapsing branches with poor support (that is, those with support lower than 10% in the Shimodaira–Hasegawa test).
Divergence time estimation
Divergence times were estimated by penalized likelihood in treePL 75 , 76 . This method is computationally efficient for datasets of this scale and typically estimates similar divergence times to more computationally intensive Bayesian analyses. The coalescent species tree topology was used as the input tree with molecular branch lengths estimated in IQ-TREE, on the basis of a concatenated alignment of the top 25 genes selected by SortaDate 77 . Genes were selected by ranking their corresponding gene trees according to the number of congruent bipartitions with the species tree. We selected genes on this basis because high gene tree conflict leads to error in divergence time estimates 78 , 79 .
Fossil calibrations were based on the AngioCal fossil calibration dataset described in ref. 5 . We used an updated version of this dataset, referred to as AngioCal v.1.1 (Supplementary Table 6 and Supplementary File 2 ). Assigning fossil calibrations in this dataset to our tree topology led to 200 unique minimum age calibrations at internal nodes (Supplementary Table 7 and Supplementary Fig. 12 ). A maximum constraint of 154 or 247 Ma was used at the angiosperm crown node. These two values, respectively, represent a young and old constraint for the maximum age of the angiosperm crown node 5 , 28 . Both values are nonetheless considerably older than the oldest known crown group angiosperm fossils of around 127.2 Ma (ref. 80 ). Both maximum constraints, in combination with all the minimum age constraints, were used to time-calibrate the species tree. Depending on the maximum constraint at the root node, these dated phylogenetic trees are referred to as young tree and old tree, respectively. For both the young tree and old tree, four analyses were performed in treePL, using smoothing values of 0.1, 1, 10 or 100. These different smoothing values assume high to low levels of among-branch substitution rate variation.
Sampling extant lineages through time
At 1 Myr intervals from the root age of the dated phylogenetic trees to the present, we calculated how many angiosperm lineages would have been present in a hypothetical tree that sampled 100% of extant angiosperm species diversity. We used this to quantify the proportion of extant lineages incorporated by our phylogenetic trees through time ( Supplementary Methods ). To do this we simulated unsampled diversity on the dated trees: the species diversity of unsampled genera was simulated as a constant-rate birth–death branching process originating in the crown group of its respective family, whilst unsampled species diversity in sampled genera was simulated as a constant-rate birth–death branching process originating at the stem node of the relevant genus. The extant diversity of each simulated branching process was determined using the World Checklist of Vascular Plants 2 . At each time interval, we then calculated the proportional difference between the number of lineages in our dated phylogenetic tree and the hypothetical fully sampled tree.
Diversification rate estimation
Dated trees estimated with alternative smoothing values were very similar (Extended Data Fig. 2 and Supplementary Fig. 5 ), so diversification rate estimates were only performed with the dated trees estimated with a smoothing value of 10. By contrast, age estimates in the young and old trees differed markedly. Diversification rate estimates were therefore performed for both these dated trees. In each case, the dated trees were pruned such that there was a maximum of one tip for each genus.
An initial analysis of diversification rates was performed by generating LTT plots as heatmaps for angiosperms as a whole, as well as for each order, with colours representing the steepness of each LTT curve at 5 Myr intervals. To calculate the steepness of the curve, we calculated the running difference between logarithmic corrected cumulative sums of lineages and applied Tukey’s running median smoothing to avoid excessive noise. For order plots, the cumulative sum starts at the first branching point, that is, order crown nodes.
Time-dependent diversification parameters (speciation and extinction rates) were also explicitly estimated across all angiosperms. These analyses were performed in RevBayes with the dnEpisodicBirthDeath function 81 . The smallest time windows in which rates were estimated were 5 Ma. However, larger windows were used toward the root of the tree such that there were at least 50 branching events in each time window. Three different models were used: equal rates of speciation and extinction across all windows; variable rates of speciation across windows but equal rates of extinction; and equal rates of speciation across windows but variable rates of extinction. Bayes factor comparison was used to compare models and offered strong support for the variable rate models but could not distinguish between the two variable rate models ( Supplementary Information ), indicating that they are probably from the same congruent set of models for the species tree 82 . In subsequent discussion we primarily refer to results from the variable speciation rate model (for justification see Supplementary Information ), although both variable rate models estimate similar patterns of net diversification rates through time ( Supplementary Information ).
Lineage-specific diversification rate estimation was performed in BAMM 83 and RevBayes. For analyses in BAMM, the setBammPriors function from the R package BAMMtools 84 was used to define appropriate priors. Different sets of analyses were performed with the prior for the expected number of shifts set to either 10 or 100. These different prior settings had minimal effect on parameter estimates. Clade-specific sampling fractions were specified for each sampled family and a backbone sampling fraction of 1 was used. We therefore accounted for incomplete sampling within families alongside comprehensive sampling of the backbone of the tree. For analyses in RevBayes, the dnCDBDP function was used and the prior for the total number of rate shifts was set to either 10 or 100. Clade-specific sampling fractions cannot be specified with this function. Therefore, the sampling fraction was set to 1 meaning that estimates will become inaccurate toward the present because of unsampled within-family diversity.
Simulations on gene tree conflict
Simulations were based on a multispecies coalescent process. Each species tree contained 100 tips and was simulated as a birth–death branching process with time-dependent rates of speciation and extinction. In experiment 1, the extinction rate was always 0. The speciation rate was 0.75 species Myr −1 at times over 6 Ma, between 6 and 2 Ma the speciation rate was 0.075 species Myr −1 and less than 2 Ma the speciation rate was 0.75 species Myr −1 . In experiment 2, the net diversification rates were the same as in experiment 1; however, in this case changes to the extinction rate led to the net diversification rate shifts. Therefore, for all time intervals, the speciation rate was 0.75 species Myr −1 . At times over 6 Ma the extinction rate was 0 species Myr −1 , between 6 and 2 Ma the extinction rate was 0.675 species Myr −1 and at times less than 2 Ma the extinction rate was 0 species Myr −1 .
Species trees with extinct lineages have extra complexities: first, changes in the extinction rate have a less direct impact on the duration of extant lineages in the species tree compared to changes in the speciation rate ( Supplementary Information ); and second, the effect of extinction is reduced at times close to the present. This causes shorter branches in the species tree, leading to the so-called ‘pull of the present’. We therefore performed a further analysis that was similar to experiment 2 but with no decrease in the extinction rate at the present. This offered insight into the effect of the ‘pull of the present’ on inferences of gene tree conflict and diversification rates and the relationship between these variables and the timing of rate shifts.
One-hundred gene trees were simulated along the branches of the birth–death branching processes according to a multispecies coalescent process. For most experiments, the effective population size was 5,000. In one further experiment, which was otherwise the same as experiment 1, the effective population size was 50,000. For each simulated dataset, the degree to which the simulated gene trees exhibited conflicting topologies with the species tree was plotted through time ( Supplementary Information ). This enabled characterization of the relationship between gene tree conflict caused by incomplete lineage sorting and shifts in speciation and extinction rates in the species tree.
More methods, results and discussion are available ( Supplementary Information ; Supplementary Figs. 13 – 24 and Supplementary Table 8 ).
Inclusion and ethics statement
The research described here results from a highly inclusive, large-scale, international collaboration, which has actively encouraged the participation of many individuals from around the world. The authorship comprises many nationalities and is representative in terms of gender, career stage and career path. A total of 163 herbaria from 48 countries provided samples and/or house herbarium vouchers related to samples used in the study (see Acknowledgements). These samples originated from many countries, including Indigenous lands. We recognize the complex histories underlying all natural history collections and the global challenge we face in acknowledging them. We gave priority to recently collected samples and, as a result, most (85%) date from the postcolonial era (estimated here as 1970 onward). To share the benefits of our research, all data generated through this collaboration have been made publicly available before the submission of this work in several data releases, starting in 2019 (see Data availability).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All raw DNA sequence data generated for this study are deposited in the European Nucleotide Archive under the following bioprojects PRJNA478314 , PRJEB35285 , PRJEB49212 and PRJNA678873 . All analysed data and metadata are available in Zenodo at https://doi.org/10.5281/zenodo.10778206 (ref. 55 ). The resulting trees and metadata are also available in GBIF ( https://doi.org/10.15468/4njn8b ) and Open Tree of Life ( https://tree.opentreeoflife.org/curator/study/view/ot_2304 ). The names used in this work match the World Checklist of Vascular Plants ( https://doi.org/10.34885/jdh2-dr22 ).
Code availability
The code used and developed to perform analyses is available in GitHub at https://github.com/RBGKew/AngiospermPhylogenomics and Zenodo at https://doi.org/10.5281/zenodo.10778206 (ref. 55 ).
Diamond, J. Evolution, consequences and future of plant and animal domestication. Nature 418 , 700–707 (2002).
Article ADS CAS PubMed Google Scholar
Govaerts, R., Nic Lughadha, E., Black, N., Turner, R. & Paton, A. The World Checklist of Vascular Plants, a continuously updated resource for exploring global plant diversity. Sci. Data 8 , 215 (2021).
Article PubMed PubMed Central Google Scholar
Chase, M. W. et al. Phylogenetics of seed plants: an analysis of nucleotide sequences from the plastid gene rbcL. Ann. Missouri. Bot. Gard. 80 , 528–580 (1993).
Article Google Scholar
Li, H.-T. et al. Plastid phylogenomic insights into relationships of all flowering plant families. BMC Biol. 19 , 232 (2021).
Ramírez-Barahona, S., Sauquet, H. & Magallón, S. The delayed and geographically heterogeneous diversification of flowering plant families. Nat. Ecol. Evol. 4 , 1232–1238 (2020).
Article PubMed Google Scholar
Li, H.-T. et al. Origin of angiosperms and the puzzle of the Jurassic gap. Nat. Plants 5 , 461–470 (2019).
Dimitrov, D. et al. Diversification of flowering plants in space and time. Nat. Commun. 14 , 7609 (2023).
Article ADS CAS PubMed PubMed Central Google Scholar
Johnson, M. G. et al. A universal probe set for targeted sequencing of 353 nuclear genes from any flowering plant designed using k-medoids clustering. Syst. Biol. 68 , 594–606 (2019).
Article CAS PubMed Google Scholar
One Thousand Plant Transcriptomes Initiative. One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574 , 679–685 (2019).
Article CAS Google Scholar
Barba-Montoya, J., dos Reis, M., Schneider, H., Donoghue, P. C. J. & Yang, Z. Constraining uncertainty in the timescale of angiosperm evolution and the veracity of a Cretaceous Terrestrial Revolution. New Phytol. 218 , 819–834 (2018).
Doyle, J. A. Molecular and fossil evidence on the origin of angiosperms. Annu. Rev. Earth Planet. Sci. 40 , 301–326 (2012).
Article ADS CAS Google Scholar
Holbourn, A. E. et al. Late Miocene climate cooling and intensification of southeast Asian winter monsoon. Nat. Commun. 9 , 1584 (2018).
Article ADS PubMed PubMed Central Google Scholar
Pennington, R. T., Cronk, Q. C. B. & Richardson, J. A. Introduction and synthesis: plant phylogeny and the origin of major biomes. Philos. Trans. R. Soc. Lond. B 359 , 1455–1464 (2004).
Benton, M. J., Wilf, P. & Sauquet, H. The Angiosperm terrestrial revolution and the origins of modern biodiversity. New Phytol. 233 , 2017–2035 (2022).
Dodsworth, S. et al. Hyb-Seq for flowering plant systematics. Trends Plant Sci. 24 , 887–891 (2019).
Brewer, G. E. et al. Factors affecting targeted sequencing of 353 nuclear genes from herbarium specimens spanning the diversity of angiosperms. Front. Plant Sci. 10 , 1102 (2019).
Baker, W. J. et al. Exploring Angiosperms353: an open, community toolkit for collaborative phylogenomic research on flowering plants. Am. J. Bot. 108 , 1059–1065 (2021).
The Angiosperm Phylogeny Group. et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181 , 1–20 (2016).
Joyce, E. M. et al. Phylogenomic analyses of Sapindales support new family relationships, rapid Mid-Cretaceous Hothouse diversification and heterogeneous histories of gene duplication. Front. Plant Sci. 14 , 1063174 (2023).
Yan, Z., Smith, M. L., Du, P., Hahn, M. W. & Nakhleh, L. Species tree inference methods intended to deal with iIncomplete lineage sorting are robust to the presence of paralogs. Syst. Biol. 71 , 367–381 (2022).
Watanabe, T., Kure, A. & Horiike, T. OrthoPhy: a program to construct ortholog data sets using taxonomic information. Genome Biol. Evol. 15 , evad026 (2023).
Ruhfel, B. R., Gitzendanner, M. A., Soltis, P. S., Soltis, D. E. & Burleigh, J. G. From algae to angiosperms—inferring the phylogeny of green plants (Viridiplantae) from 360 plastid genomes. BMC Evol. Biol. 14 , 23 (2014).
Endress, P. K. Origins of flower morphology. J. Exp. Zool. 291 , 105–115 (2001).
Stull, G. W., Duno de Stefano, R., Soltis, D. E. & Soltis, P. S. Resolving basal lamiid phylogeny and the circumscription of Icacinaceae with a plastome-scale data set. Am. J. Bot. 102 , 1794–1813 (2015).
Soltis, D. E. et al. Chloroplast gene sequence data suggest a single origin of the predisposition for symbiotic nitrogen fixation in angiosperms. Proc. Natl Acad. Sci. USA 92 , 2647–2651 (1995).
Walker, J. F. et al. From cacti to carnivores: improved phylotranscriptomic sampling and hierarchical homology inference provide further insight into the evolution of Caryophyllales. Am. J. Bot. 105 , 446–462 (2018).
Guo, X. et al. Chloranthus genome provides insights into the early diversification of angiosperms. Nat. Commun. 12 , 6930 (2021).
Sauquet, H., Ramírez-Barahona, S. & Magallón, S. What is the age of flowering plants? J. Exp. Bot. 73 , 3840–3853 (2022).
Landis, J. B. et al. Impact of whole-genome duplication events on diversification rates in angiosperms. Am. J. Bot. 105 , 348–363 (2018).
Tank, D. C. et al. Nested radiations and the pulse of angiosperm diversification: increased diversification rates often follow whole genome duplications. New Phytol. 207 , 454–467 (2015).
Darwin, C. The Correspondence of Charles Darwin (Cambridge Univ. Press, 1879).
Buggs, R. J. A. Reconfiguring Darwin’s abominable mystery. Nat. Plants 8 , 194–195 (2022).
Coiro, M., Doyle, J. A. & Hilton, J. How deep is the conflict between molecular and fossil evidence on the age of angiosperms? New Phytol. 223 , 83–99 (2019).
Herendeen, P. S., Friis, E. M., Pedersen, K. R. & Crane, P. R. Palaeobotanical redux: revisiting the age of the angiosperms. Nat. Plants 3 , 17015 (2017).
Friis, E. M., Crane, P. R. & Pedersen, K. R. Early Flowers and Angiosperm Evolution (Cambridge Univ. Press, 2011).
Davies, T. J. et al. Darwin’s abominable mystery: insights from a supertree of the angiosperms. Proc. Natl Acad. Sci. USA 101 , 1904–1909 (2004).
Magallón, S., Gómez-Acevedo, S., Sánchez-Reyes, L. L. & Hernández-Hernández, T. A metacalibrated time-tree documents the early rise of flowering plant phylogenetic diversity. New Phytol. 207 , 437–453 (2015).
Mathews, S. & Donoghue, M. J. The root of angiosperm phylogeny inferred from duplicate phytochrome genes. Science 286 , 947–950 (1999).
Dilcher, D. Toward a new synthesis: major evolutionary trends in the angiosperm fossil record. Proc. Natl Acad. Sci. USA 97 , 7030–7036 (2000).
Meredith, R. W. et al. Impacts of the Cretaceous terrestrial revolution and KPg extinction on mammal diversification. Science 334 , 521–524 (2011).
Bouchenak-Khelladi, Y., Onstein, R. E., Xing, Y., Schwery, O. & Linder, H. P. On the complexity of triggering evolutionary radiations. New Phytol. 207 , 313–326 (2015).
Donoghue, M. J. & Sanderson, M. J. Confluence, synnovation and depauperons in plant diversification. New Phytol. 207 , 260–274 (2015).
Magallón, S., Sánchez-Reyes, L. L. & Gómez-Acevedo, S. L. Thirty clues to the exceptional diversification of flowering plants. Ann. Bot. 123 , 491–503 (2019).
Rabosky, D. L. Diversity-dependence, ecological speciation and the role of competition in macroevolution. Annu. Rev. Ecol. Evol. Syst. 44 , 481–502 (2013).
Asar, Y., Ho, S. Y. W. & Sauquet, H. Early diversifications of angiosperms and their insect pollinators: were they unlinked? Trends Plant Sci. 27 , 858–869 (2022).
Peris, D. & Condamine, F. L. The dual role of the angiosperm radiation on insect diversification. Nat. Commun. 15 , 552 (2024).
Folk, R. A. et al. Rates of niche and phenotype evolution lag behind diversification in a temperate radiation. Proc. Natl Acad. Sci. USA 116 , 10874–10882 (2019).
Sun, M. et al. Recent accelerated diversification in rosids occurred outside the tropics. Nat. Commun. 11 , 3333 (2020).
Soltis, P. S., Folk, R. A. & Soltis, D. E. Darwin review: angiosperm phylogeny and evolutionary radiations. Proc. R. Soc. B 286 , 20190099 (2019).
Article PubMed Central Google Scholar
Baker, W. J. et al. A comprehensive phylogenomic platform for exploring the angiosperm tree of life. Syst. Biol. 71 , 301–319 (2022).
McDonnell, A. J. et al. Exploring Angiosperms353: developing and applying a universal toolkit for flowering plant phylogenomics. Appl. Plant Sci. 9 , e11443 (2021).
Bratzel, F. et al. Target-enrichment sequencing reveals for the first time a well-resolved phylogeny of the core Bromelioideae (family Bromeliaceae). TAXON 72 , 47–63 (2023).
Gagnon, E. et al. Phylogenomic discordance suggests polytomies along the backbone of the large genus Solanum. Am. J. Bot. 109 , 580–601 (2022).
Murillo-A, J., Valencia-D, J., Orozco, C. I., Parra-O, C. & Neubig, K. M. Incomplete lineage sorting and reticulate evolution mask species relationships in Brunelliaceae, an Andean family with rapid, recent diversification. Am. J. Bot. 109 , 1139–1156 (2022).
Zuntini, A.R. & Carruthers, T. Phylogenomics and the rise of the angiosperms. Zenodo https://doi.org/10.5281/zenodo.10778206 (2024).
Hendriks, K. P. et al. Global Brassicaceae phylogeny based on filtering of 1,000-gene dataset. Curr. Biol 33 , 4052–4068 (2023).
Chen, L.-Y. et al. Phylogenomic analyses of Alismatales shed light into adaptations to aquatic environments. Mol. Biol. Evol. 39 , msac079 (2022).
Article CAS PubMed PubMed Central Google Scholar
Timilsena, P. R. et al. Phylogenomic resolution of order- and family-level monocot relationships using 602 single-copy nuclear genes and 1375 BUSCO genes. Front. Plant Sci. 13 , 876779 (2022).
Ogutcen, E. et al. Phylogenomics of Gesneriaceae using targeted capture of nuclear genes. Mol. Phylogenet. Evol. 157 , 107068 (2021).
Yardeni, G. et al. Taxon-specific or universal? Using target capture to study the evolutionary history of rapid radiations. Mol. Ecol. Resour. 22 , 927–945 (2022).
Johnson, M. G. et al. HybPiper: extracting coding sequence and introns for phylogenetics from high-throughput sequencing reads using target enrichment. Appl. Plant Sci. 4 , 1600016 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30 , 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19 , 455–477 (2012).
Article MathSciNet CAS PubMed PubMed Central Google Scholar
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6 , 31 (2005).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: improvements in performance and usability. Mol. Biol. Evol. 30 , 772–780 (2013).
Smith, S. A. & Dunn, C. W. Phyutility: a phyloinformatics tool for trees, alignments and molecular data. Bioinformatics 24 , 715–716 (2008).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37 , 1530–1534 (2020).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35 , 518–522 (2018).
Mai, U. & Mirarab, S. TreeShrink: fast and accurate detection of outlier long branches in collections of phylogenetic trees. BMC Genomics 19 , 272 (2018).
Zhang, C., Rabiee, M., Sayyari, E. & Mirarab, S. ASTRAL-III: polynomial time species tree reconstruction from partially resolved gene trees. BMC Bioinformatics 19 , 153 (2018).
Junier, T. & Zdobnov, E. M. The Newick utilities: high-throughput phylogenetic tree processing in the Unix shell. Bioinformatics 26 , 1669–1670 (2010).
Borowiec, M. L. AMAS: a fast tool for alignment manipulation and computing of summary statistics. PeerJ 4 , e1660 (2016).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5 , e9490 (2010).
Yin, J., Zhang, C. & Mirarab, S. ASTRAL-MP: scaling ASTRAL to very large datasets using randomization and parallelization. Bioinformatics 35 , 3961–3969 (2019).
Sanderson, M. J. Estimating absolute rates of molecular evolution and divergence times: a penalized likelihood approach. Mol. Biol. Evol. 19 , 101–109 (2002).
Smith, S. A. & O’Meara, B. C. treePL: divergence time estimation using penalized likelihood for large phylogenies. Bioinformatics 28 , 2689–2690 (2012).
Smith, S. A., Brown, J. W. & Walker, J. F. So many genes, so little time: a practical approach to divergence-time estimation in the genomic era. PLoS ONE 13 , e0197433 (2018).
Britton, T. Estimating divergence times in phylogenetic trees without a molecular vlock. Syst. Biol. 54 , 500–507 (2005).
Carruthers, T. et al. The implications of incongruence between gene tree and species tree topologies for divergence time estimation. Syst. Biol. 71 , 1124–1146 (2022).
Gomez, B., Daviero-Gomez, V., Coiffard, C., Martín-Closas, C. & Dilcher, D. L. Montsechia, an ancient aquatic angiosperm. Proc. Natl Acad. Sci. USA 112 , 10985–10988 (2015).
Höhna, S. et al. RevBayes: Bayesian phylogenetic inference using graphical models and an interactive model-specification language. Syst. Biol. 65 , 726–736 (2016).
Louca, S. & Pennell, M. W. Extant timetrees are consistent with a myriad of diversification histories. Nature 580 , 502–505 (2020).
Rabosky, D. L. Automatic detection of key innovations, rate shifts and diversity-dependence on phylogenetic trees. PLoS ONE 9 , e89543 (2014).
Rabosky, D. L. et al. BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees. Methods Ecol. Evol. 5 , 701–707 (2014).
Download references
Acknowledgements
The PAFTOL project was funded by grants from the Calleva Foundation to the Royal Botanic Gardens, Kew. Data were also contributed by the Genomics for Australian Plants Framework Initiative consortium funded by Bioplatforms Australia (enabled by the National Collaborative Research Infrastructure Strategy) and partner organizations. The work was further supported by research grants from VILLUM FONDEN (grant no. 00025354) and the Aarhus University Research Foundation (grant no. AUFF-E-2017-7-19) to W.L.E. and from grant nos NSF DBI 1930030 and DEB 1917146 to S.A.S. Computational resources and technical support were provided by the Research/Scientific Computing teams at The James Hutton Institute and the National Institute of Agricultural Botany (NIAB) through the ‘UK’s Crop Diversity Bioinformatics HPC’ (BBSRC grant no. BB/S019669/1). The following provided technical assistance to the project at various stages: O. Berry, N. Black, M. Corcoran, S. Dequiret, I. Fairlie, L. Frankel, T. Freeth, A. Gilbert, B. Lepschi, D. Lewis, L. May, A. McArdle, E. O’Loughlin, S. Phillips, T. Sarkinen, L. Simmons, N. Walsh and M.-H. Weech. We thank all institutions who made their biological collections available and the many botanists and co-workers in the field who have collected, identified and curated the specimens used in this project. Specifically, we thank the following herbaria and their staff for providing samples for genomic analysis and/or for housing voucher specimens associated with analysed samples: A, ABH, AD, ALTB, APSC, B, BA, BC, BCN, BCRU, BG, BH, BHCB, BISH, BJFC, BKF, BM, BNRH, BOL, BONN, BR, BRI, BRIT, BRLU, BRUN, C, CAN, CANB, CAS, CBG, CNS, COL, CONC, CORD, CS, CTES, CUVC, DNA, E, EA, F, FI, FLAS, FMB, FTG, G, GB, GC, GENT, GH, GOET, GUAY, GZU, HAW, HEID, HITBC, HNG, HO, HPUJ, HRCB, HTW, HUA, HUAL, HUAZ, HUB, HUEFS, HUFU, IBSC, IBUG, ICN, IEB, INB, INPA, JBB, JBL, JRAU, K, KAS, KLU, KRB, KUN, L, LE, LISC, LP, LPB, LYJB, M, MA, MAU, MBA, MBML, MEDEL, MEL, MELU, MHA, MICH, MIN, MJG, MO, MSUN, MT, MY, N, NBG, NCU, NCY, NE, NH, NHM, NHMR, NMNL, NOU, NSW, NU, NY, OS, OSBU, P, PERTH, PG, PH, PRE, PTBG, QBG, QCA, QRS, RB, REU, S, SALA, SAR, SEV, SGO, SI, SING, SP, SPF, SPFR, SUVA, TCD, TEX, TNS, TUH, TUM, U, UAPC, UB, UBT, UDW, UEC, UPCB, UPR, UPS, UPTC, US, USM, W, WAG, WS, WTU, YA and ZSS; acronyms follow Index Herbariorum ( https://sweetgum.nybg.org/science/ih/ ). We also thank the Millennium Seed Bank Partnership for supporting access to samples. We acknowledge all national, state and regional authorities who authorized and facilitated the sourcing of these specimens. See also extended acknowledgements in the Supplementary Information .
Author information
These authors contributed equally: Alexandre R. Zuntini, Tom Carruthers
These authors jointly supervised this work: Stephen A. Smith, Wolf L. Eiserhardt, Félix Forest, William J. Baker
Authors and Affiliations
Royal Botanic Gardens, Kew, Richmond, UK
Alexandre R. Zuntini, Tom Carruthers, Olivier Maurin, Paul C. Bailey, Kevin Leempoel, Grace E. Brewer, Niroshini Epitawalage, Elaine Françoso, Berta Gallego-Paramo, Catherine McGinnie, Raquel Negrão, Shyamali R. Roy, Eduardo Toledo Romero, Vanessa M. A. Barber, James J. Clarkson, Robyn S. Cowan, Lisa Pokorny, Ai-Qun Hu, Isabel Larridon, Panagiota Malakasi, Natalia A. S. Przelomska, Toral Shah, Juan Viruel, Watchara Arthan, Nicky Biggs, Renata Borosova, Gemma L. C. Bramley, Marie Briggs, Edie Burns, Stuart Cable, Abigail J. A. Carruthers, Mark W. Chase, Martin Cheek, Maarten J. M. Christenhusz, Laszlo Csiba, Iain Darbyshire, Nina M. J. Davies, Aaron P. Davis, Sara L. Edwards, Michael F. Fay, Sarah Z. Ficinski, Sue Frisby, Tim Fulcher, David J. Goyder, Aurélie Grall, Laura Green, Jan Hackel, Anna Haigh, Tony Hall, Sebastian A. Hatt, Helen C. F. Hopkins, Imalka M. Kahandawala, Bente B. Klitgaard, Christine J. Leon, Gwilym P. Lewis, Meng Lu, Eve J. Lucas, Manuel Luján, Carlos Magdalena, Lizo E. Masters, Simon J. Mayo, Alexandre K. Monro, Oscar A. Perez-Escobar, Robyn F. Powell, Ghillean T. Prance, Carmen Puglisi, Paul E. J. Rees, Saba Rokni, Ana Rita G. Simões, Michelle Siros, Cynthia A. Sothers, Anna Trias-Blasi, Timothy M. A. Utteridge, Maria S. Vorontsova, Noor Al-Wattar, Roseina Woods, Martin Xanthos, Sue Zmarzty, Alexandre Antonelli, Sidonie Bellot, Olwen M. Grace, Paul J. Kersey, Ilia J. Leitch, Wolf L. Eiserhardt, Félix Forest & William J. Baker
Centre for Ecology, Evolution and Behaviour, Department of Biological Sciences, School of Life Sciences and the Environment, Royal Holloway University of London, London, UK
Elaine Françoso
Australian Tropical Herbarium, James Cook University, Smithfield, Queensland, Australia
Lalita Simpson, Elizabeth M. Joyce, Matthew D. Barrett, Melissa J. Harrison, Katharina Nargar, Lars Nauheimer, Stuart Worboys & Darren M. Crayn
Centre for Research in Agricultural Genomics (CRAG), CSIC-IRTA-UAB-UB, Campus UAB, Barcelona, Spain
Laura Botigué
School of Biological Sciences, University of Portsmouth, Portsmouth, UK
Steven Dodsworth & Natalia A. S. Przelomska
Texas Tech University, Lubbock, TX, USA
Matthew G. Johnson
School of Physics, Engineering and Computer Science, University of Hertfordshire, Hatfield, UK
Department of Biodiversity and Conservation, Real Jardín Botánico (RJB-CSIC), Madrid, Spain
Lisa Pokorny
Department of Biological Sciences, Clemson University, Clemson, SC, USA
Norman J. Wickett
Departamento de Botânica, Instituto de Biociências, Universidade de São Paulo, São Paulo, Brazil
Guilherme M. Antar, Elton John de Lírio & José R. Pirani
Departamento de Ciências Agrárias e Biológicas, Centro Universitário Norte do Espírito Santo, Universidade Federal do Espírito Santo, São Mateus, Brazil
Guilherme M. Antar
Smith College, Northampton, MA, USA
Lucinda DeBolt & Karime Gutierrez
Department of Biology, University of Osnabrück, Osnabrück, Germany
Kasper P. Hendriks & Klaus Mummenhoff
Naturalis Biodiversity Center, Leiden, The Netherlands
Kasper P. Hendriks, Diego Bogarín, Roy H. J. Erkens, Frederic Lens, Vincent S. F. T. Merckx & Jan J. Wieringa
Plant Biodiversity, Technical University Munich, Freising, Germany
Alina Hoewener, Edgardo M. Ortiz & Hanno Schaefer
Systematic, Biodiversity and Evolution of Plants, Ludwig Maximilian University of Munich, Munich, Germany
Elizabeth M. Joyce
Department of Botany, University of British Columbia, Vancouver, British Columbia, Canada
Izai A. B. S. Kikuchi & Sean W. Graham
Department of Ecology & Evolutionary Biology, University of Michigan, Ann Arbor, MI, USA
Drew A. Larson, Richard K. Rabeler & Stephen A. Smith
Germplasm Bank of Wild Species, Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, China
Jing-Xia Liu, De-Zhu Li & Meng-Yuan Zhou
Royal Botanic Gardens Victoria, Melbourne, Victoria, Australia
Theodore R. Allnutt, Helen F. Barnes, David J. Cantrill, Bee F. Gunn, Gareth D. Holmes, Christopher J. Jackson, Todd G. B. McLay, Daniel J. Murphy & Frank Udovicic
Department of Plant and Environmental Biology, University of Ghana, Accra, Ghana
Gabriel K. Ameka
Botany and N.C.W. Beadle Herbarium, University of New England, Armidale, New South Wales, Australia
Rose L. Andrew, Jeremy J. Bruhl, Ian R. H. Telford & Andrew H. Thornhill
Department of Systematics, Biodiversity and Evolution of Plants, Albrecht-von-Haller Institute of Plant Sciences, University of Göttingen, Göttingen, Germany
Marc S. Appelhans
Departamento de Biología Vegetal y Ecología, Facultad de Biología, Universidad de Sevilla, Seville, Spain
Montserrat Arista, Juan Arroyo, Alejandra de Castro Mateo, Marcial Escudero & Jose C. Del Valle
General Research Services, Herbario SEV, CITIUS, Universidad de Sevilla, Seville, Spain
María Jesús Ariza
Institute of Biology, Freie Universität, Berlin, Germany
Julien B. Bachelier
Department of Biology, New Mexico State University, Las Cruces, NM, USA
C. Donovan Bailey
National Herbarium of NSW, Botanic Gardens of Sydney, Mount Annan, New South Wales, Australia
Russell L. Barrett, Marco F. Duretto, Richard W. Jobson, Patricia Lu-Irving, Kristina McColl, Hannah McPherson, Matthew Renner, Ifeanna Tooth, Trevor C. Wilson, Lisa A. Woods & Hervé Sauquet
Department of Biological Sciences, University of Memphis, Memphis, TN, USA
Randall J. Bayer
School of BioSciences, The University of Melbourne, Parkville, Victoria, Australia
Michael J. Bayly, Joanne L. Birch & Rachael M. Fowler
State Herbarium of South Australia, Botanic Gardens and State Herbarium, Adelaide, South Australia, Australia
Ed Biffin, Ainsley Calladine, Francis J. Nge, Andrew H. Thornhill, Helen P. Vonow & Michelle Waycott
Jardín Botánico Lankester, Universidad de Costa Rica, Cartago, Costa Rica
Diego Bogarín
School of Geographical Sciences, University of Bristol, Bristol, UK
Alexander M. C. Bowles
Centro Studi Erbario Tropicale, Dipartimento di Biologia, University of Florence, Florence, Italy
Peter C. Boyce
Centre for Australian National Biodiversity Research, National Research Collections Australia, CSIRO, Canberra, Australian Capital Territory, Australia
Linda Broadhurst, Mark A. Clements, Katharina Nargar & Alexander Schmidt-Lebuhn
Queensland Herbarium and Biodiversity Science, Brisbane Botanic Gardens, Toowong, Queensland, Australia
Gillian K. Brown
Institut de Recherche en Biologie Végétale and Département de Sciences Biologiques, University of Montreal, Montreal, Quebec, Canada
Anne Bruneau
Department of Biological Sciences, Boise State University, Boise, ID, USA
Sven Buerki & James F. Smith
Biodiversity and Conservation Science, Department of Biodiversity, Conservation and Attractions, Government of Western Australia, Kensington, Western Australia, Australia
Margaret Byrne
Conservatoire et Jardin Botaniques de Genève, Chambésy, Switzerland
Martin W. Callmander
Cambridge University Botanic Garden, Cambridge, UK
Ángela Cano
Department of Forest and Conservation Sciences, University of British Columbia, Vancouver, British Columbia, Canada
Warren M. Cardinal-McTeague
Missouri Botanical Garden, St. Louis, MO, USA
Mónica M. Carlsen, Gerrit Davidse, Carmen Puglisi, Ihsan Al-Shehbaz & Peter F. Stevens
Department of Environment and Agriculture, Curtin University, Bentley, Western Australia, Australia
Mark W. Chase
Department of Biology, Ghent University, Ghent, Belgium
Lars W. Chatrou & Federico Fabriani
Institute of Herbgenomics, Chengdu University of Traditional Chinese Medicine, Chengdu, China
Shilin Chen
Institute of Medicinal Plant Development, Chinese Academy of Medical Sciences, Beijing, China
Department of Environment and Agriculture, Curtin University, Perth, Western Australia, Australia
Maarten J. M. Christenhusz
Plant Gateway, Den Haag, The Netherlands
Ecology and Evolutionary Biology, School of Biosciences, University of Sheffield, Sheffield, UK
Pascal-Antoine Christin
Western Australian Herbarium, Department of Biodiversity, Conservation and Attractions, Government of Western Australia, Kensington, Western Australia, Australia
Skye C. Coffey, Shelley A. James, Terry D. Macfarlane & Kelly A. Shepherd
School of Biological Sciences, The University of Adelaide, Adelaide, South Australia, Australia
John G. Conran, Andrew H. Thornhill & Michelle Waycott
Herbario GUAY, Facultad de Ciencias Naturales, Universidad de Guayaquil, Guayaquil, Ecuador
Xavier Cornejo
DIADE, Université Montpellier, CIRAD IRD, Montpellier, France
Thomas L. P. Couvreur
Northern Territory Herbarium Department of Environment Parks & Water Security, Northern Territory Government, Palmerston, Northern Territory, Australia
Ian D. Cowie
The University of Adelaide, North Terrace Campus, Adelaide, South Australia, Australia
Kor-jent van Dijk, Andrew E. McDougall & Luis T. Williamson
Department of Plant Biology, University of Illinois at Urbana-Champaign, Urbana, IL, USA
Stephen R. Downie
Department of Biological Sciences and Institute for the Study of the Environment, Sustainability and Energy, Northern Illinois University, DeKalb, IL, USA
Melvin R. Duvall
Sukkulenten-Sammlung Zürich/ Grün Stadt Zürich, Zürich, Switzerland
Maastricht Science Programme, Maastricht University, Maastricht, The Netherlands
Roy H. J. Erkens
System Earth Science, Maastricht University, Venlo, The Netherlands
Departamento de Botánica, Ecología y Fisiología Vegetal, Facultad de Ciencias, Universidad de Córdoba, Córdoba, Spain
Manuel de la Estrella
Departamento de Biologia, Faculdade de Ciências e Letras de Ribeirão Preto, Universidade de São Paulo, São Paulo, Brazil
Paola de L. Ferreira
Department of Biology, Aarhus University, Aarhus, Denmark
Paola de L. Ferreira, Wolf L. Eiserhardt & William J. Baker
South China Botanical Garden, Chinese Academy of Sciences, Guangzhou, China
Lin Fu & Ming Qin
Systematics and Evolution of Vascular Plants (UAB)—Associated Unit to CSIC by IBB, Departament de Biologia Animal, Biologia Vegetal i Ecologia, Facultat de Biociències, Universitat Autònoma de Barcelona, Bellaterra, Spain
Mercè Galbany-Casals
Department of Biology, Case Western Reserve University, Cleveland, OH, USA
Elliot M. Gardner
Altai State University, Barnaul, Russia
Dmitry A. German
Faculdade de Ciências Biológicas e Ambientais, Universidade Federal da Grande Dourados, Dourados, Brazil
Augusto Giaretta
Laboratoire Sciences Pour l’Environnement, Université de Corse, Ajaccio, France
Marc Gibernau
Canadian Museum of Nature, Ottawa, Ontario, Canada
Lynn J. Gillespie
Herbario Trelew, Universidad Nacional de la Patagonia San Juan Bosco, Trelew, Argentina
Cynthia C. González
Museo Argentino de Ciencias Naturales (MACN-CONICET), Buenos Aires, Argentina
Diego G. Gutiérrez & Luis Palazzesi
Department of Biology, Universität Marburg, Marburg, Germany
Institut de Systématique, Evolution, Biodiversité, Muséum National d’Histoire Naturelle, Paris, France
Thomas Haevermans & Marc Pignal
Department of Biological Sciences, University of Alberta, Edmonton, Alberta, Canada
Jocelyn C. Hall
Institut Botànic de Barcelona (IBB CSIC-Ajuntament de Barcelona), Barcelona, Spain
Oriane Hidalgo & Jaume Pellicer
Botany, School of Natural Sciences, Trinity College Dublin, The University of Dublin, Dublin, Ireland
Trevor R. Hodkinson
Prinzessin Therese von Bayern-Lehrstuhl für Systematik, Biodiversität & Evolution der Pflanzen, Ludwig-Maximilians-Universität München, Botanische Staatssammlung München, Botanischer Garten München-Nymphenburg, Munich, Germany
Gudrun Kadereit
Gothenburg Botanical Garden, Gothenburg, Sweden
Kent Kainulainen
National Museum of Nature and Science, Tsukuba, Japan
Masahiro Kato
Donald Danforth Plant Science Center, St. Louis, MO, USA
Elizabeth A. Kellogg
Southern Cross University, Lismore, New South Wales, Australia
Graham J. King
Synergy SRG, Luton, UK
Beata Klejevskaja
Foundational Biodiversity Science Division, South African National Biodiversity Institute, Pretoria, South Africa
Ronell R. Klopper
Department of Plant and Soil Sciences, University of Pretoria, Pretoria, South Africa
Natural History Museum, London, UK
Sandra Knapp
Centre for Organismal Studies, Biodiversity and Plant Systematics, Heidelberg University, Heidelberg, Germany
Marcus A. Koch
Department of Plant Biology, University of Georgia, Athens, GA, USA
James H. Leebens-Mack
Institut de Recherche en Biologie Végétale, University of Montreal, Montreal, Quebec, Canada
Étienne Léveillé-Bourret
CSIRO, Canberra, Australian Capital Territory, Australia
Lan Li & Jennifer M. Taylor
Department of Plant Systematics, University of Bayreuth, Bayreuth, Germany
Sigrid Liede-Schumann
Department of Biodiversity, Earth and Environmental Sciences, Drexel University, Philadelphia, PA, USA
Tatyana Livshultz
Academy of Natural Science, Drexel University, Philadelphia, PA, USA
National Tropical Botanical Garden, Kalaheo, HI, USA
David Lorence
Instituto de Pesquisas Jardim Botânico do Rio de Janeiro, Rio de Janeiro, Brazil
Jaquelini Luber, Vidal F. Mansano & Ariane Luna Peixoto
Bioplatforms Australia Ltd, Sydney, New South Wales, Australia
Department of Biological Sciences, Saint Cloud State University, Saint Cloud, MN, USA
Angela J. McDonnell
Instituto de Arqueología y Antropología, Universidad Católica del Norte, San Pedro de Atacama, Chile
Rosa I. Meneses
New York Botanical Garden, Bronx, NY, USA
Fabián A. Michelangeli, John D. Mitchell & Gregory M. Plunkett
Department of Biology, Oberlin College, Oberlin, OH, USA
Michael J. Moore
Department of Ecology, Evolution & Behavior, University of Minnesota, St. Paul, MN, USA
Taryn L. Mueller
AMAP Lab, Université Montpellier, IRD, CIRAD, CNRS INRAE, Montpellier, France
Jérôme Munzinger
Laboratorio de Ecofisiología, Escuela de Ciencias Biológicas, Pontificia Universidad Católica del Ecuador, Quito, Ecuador
Priscilla Muriel
Department of Systematic and Evolutionary Botany, University of Zürich, Zürich, Switzerland
Reto Nyffeler & Rolf Rutishauser
Royal Botanic Garden Edinburgh, Edinburgh, UK
Andrés Orejuela & Olwen M. Grace
Grupo de Investigación en Recursos Naturales Amazónicos, Instituto Tecnológico del Putumayo, Mocoa, Colombia
Andrés Orejuela
US Botanic Garden, Washington, DC, USA
Susan K. Pell
Department of Biology and Marine Biology, University of North Carolina Wilmington, Wilmington, NC, USA
Darin S. Penneys
Department of Biological and Environmental Sciences, University of Gothenburg, Gothenburg, Sweden
Claes Persson & Alexandre Antonelli
LSTM Université Montpellier, CIRADIRD, Montpellier, France
Yohan Pillon
School of Biological Sciences, Washington State University, Pullman, WA, USA
Eric H. Roalson
National Parks Board, Singapore Botanic Gardens, Singapore, Singapore
Michele Rodda
New Mexico State University, Las Cruces, NM, USA
Zachary S. Rogers
Tasmanian Herbarium, University of Tasmania, Sandy Bay, Tasmania, Australia
Miguel F. de Salas
University of Exeter, Exeter, UK
Rowan J. Schley
School of Science Technology and Engineering, Center for Bioinnovation, University Sunshine Coast, Sippy Downs, Queensland, Australia
Alison Shapcott
Department of Biology, Colorado State University, Fort Collins, CO, USA
Mark P. Simmons
Departamento de Biologia Vegetal, Universidade Estadual de Campinas, Campinas, Brazil
André O. Simões
University of California, San Francisco, San Francisco, CA, USA
Michelle Siros
Departamento de Botânica, Universidade Federal do Paraná, Curitiba, Brazil
Eric C. Smidt
Pittsburg State University, Pittsburg, KS, USA
Florida Museum of Natural History, University of Florida, Gainesville, FL, USA
Douglas E. Soltis & Pamela S. Soltis
Smithsonian Institution, Washington, DC, USA
Robert J. Soreng
Department of Biology, University of Ottawa, Ottawa, Ontario, Canada
Julian R. Starr
Hobart and William Smith Colleges, Geneva, NY, USA
Shannon C. K. Straub
Rutgers University, New Brunswick, NJ, USA
Lena Struwe
Department of Biological Sciences and Bolus Herbarium, University of Cape Town, Cape Town, South Africa
G. Anthony Verboom
Department of Environmental Sciences—Botany, University of Basel, Basel, Switzerland
Jurriaan M. de Vos
Instituto de Biologia, Universidade Federal de Uberlândia, Uberlândia, Brazil
Cassiano A. D. Welker
Australian National Herbarium, Centre for Australian National Biodiversity Research, National Research Collections Australia, CSIRO, Canberra, Australian Capital Territory, Australia
Adam J. White
Institute of Biodiversity And Environmental Conservation, Universiti Malaysia Sarawak, Samarahan, Malaysia
Sin Yeng Wong
University of Minnesota-Twin Cities, St. Paul, MN, USA
Southwest Forestry University, Kunming, China
Yu-Xiao Zhang
Instituto de Botánica Darwinion, San Isidro, Argentina
Fernando O. Zuloaga
Gothenburg Global Biodiversity Centre, University of Gothenburg, Gothenburg, Sweden
Alexandre Antonelli
Department of Biology, University of Oxford, Oxford, UK
You can also search for this author in PubMed Google Scholar
Contributions
A.R.Z., T.C., A.A., S. Bellot, D.M.C., O.M.G., P.J.K., I.J.L., H. Sauquet, S.A.S., W.L.E., F. Forest and W.J.B were involved in conceptualization of this work. A.R.Z., T.C., A.A., S. Bellot, D.M.C., O.M.G., H. Sauquet, S.A.S., W.L.E., F. Forest and W.J.B. contributed to the methodology. A.R.Z., O.M., E.F., C. McGinnie, S.R.R., L. Simpson, J.J.C., R.S.C., S.D., L. Pokorny, G.M.A., K.G., K.P.H., A. Hoewener, A.-Q.H., E.M.J., I.A.B.S.K., I.L., D.A.L., E. J. Lírio, J.-X.L., P. Malakasi, N.A.S.P., T.S., J.V., G.K.A., R.L.A., M.S.A., M.A., M.J.A., J.A., W.A., J.B.B., C.D.B., H.F.B., M.D.B., R.L.B., R.J.B., M.J.B., E. Biffin, N.B., J.L.B., D.B., R.B., A.M.C.B., P. C. Boyce, G.L.C.B., M. Briggs, L. Broadhurst, G.K.B., J.J.B., A.B., S. Buerki, E. Burns, M. Byrne, S. Cable, A.C., M. W. Callmander, Á.C., D.J.C., W.M.C.-M., M.M.C., A.J.A.C., A.C.M., M. W. Chase, L.W.C., M.C., S. Chen, M.J.M.C., P.-A.C., M.A.C., S.C.C., J.G.C., X.C., T.L.P.C., I.D.C., L.C., I.D., G.D., N.M.J.D., A.P.D., K.-J.D., S.R.D., M.F.D., M.R.D., S.L.E., U.E., R.H.J.E., M. Escudero, M. Estrella, F. Fabriani, M.F.F., P.L.F., S.Z.F., R.M.F., S.F., L.F., T.F., M.G.-C., E.M.G., D.A.G., A. Giaretta, M.G., L.J.G., C.C.G., D.J.G., S.W.G., A. Grall, L.G., B.F.G., D.G.G., J.H., T. Haevermans, A. Haigh, J.C.H., T. Hall, M.J.H., S.A.H., O.H., T.R.H., G.D.H., H.C.F.H., C.J.J., S.A.J., R.W.J., G.K., I.M.K., K.K., M.K., E.A.K., G.J.K., B.K., B.B.K., R.R.K., S.K., M.A.K., J.H.L.-M., F.L., C.J.L., É.L.-B., G.P.L., D.-Z.L., L.L., S.L.-S., T.L., D.L., M. Lu, P.L.-I., J.L., E. J. Lucas, M. Luján, M. Lum, T.D.M., C. Magdalena, V.F.M., L.E.M., S.J.M., K. McColl, A.J.M., A.E.M., T.G.B.M., H.M., R.I.M., V.S.F.T.M., F.A.M., J.D.M., A.K.M., M.J.M., T.L.M., K. Mummenhoff, J.M., P. Muriel, D.J.M., K.N., L.N., F.J.N., R. Nyffeler, A.O., E.M.O., L. Palazzesi, A.L.P., S.K.P., J.P., D.S.P., O.A.P.-E., C. Persson, M.P., Y.P., J.R.P., G.M.P., R.F.P., G.T.P., C. Puglisi, M.Q., R.K.R., P.E.J.R., M. Renner, E.H.R., M. Rodda, Z.S.R., S.R., R.R., M.F.S., H. Schaefer, R. J. Schley, A.S.-L., A.S., I.S., K.A.S., M.P.S., A.O.S., A.R.G.S., M.S., E.C.S., J.F.S., N.S., D.E.S., P.S.S., R. J. Soreng, C.A.S., J.R.S., P.F.S., S.C.K.S., L. Struwe, J.M.T., I.R.H.T., A.H.T., I.T., A.T.-B., F.U., T.M.A.U., J.C.V., G.A.V., H.P.V., M.S.V., J.M.V., N.W., M.W., C.A.D.W., A.J.W., J.J.W., L.T.W., T.C.W., S.Y.W., L.A.W., R.W., S.W., M.X., Y.Y., Y.-X.Z., M.-Y.Z., S.Z., F.O.Z., S. Bellot, D.M.C., O.M.G., H. Sauquet, W.L.E., F. Forest and W.J.B. provided resources. A.R.Z., T.C., O.M., P. C. Bailey, K.L., G.E.B., N.E., E.F., B.G.-P., C. McGinnie, S.R.R., L. Simpson, L. Botigué, J.J.C., R.S.C., S.D., M.G.J., J.T.K., L. Pokorny, N.J.W., G.M.A., L.D., K.G., K.P.H., A. Hoewener, A.-Q.H., E.M.J., I.A.B.S.K., I.L., D.A.L., E. J. Lírio, J.-X.L., P. Malakasi, N.A.S.P., T.S., J.V. and S. Bellot carried out the investigations. A.R.Z., T.C., O.M., G.E.B., N.E., E.F., B.G.-P., C. McGinnie, R. Negrão, S.R.R., L. Simpson, E.T.R., V.M.A.B., K.P.H., J.V., T.R.A. and H. Sauquet were responsible for data curation. A.R.Z., T.C., P. C. Bailey and K.L. conducted the formal analysis. A.R.Z., T.C., P. C. Bailey, K.L., M.G.J. and J.T.K. developed the software. A.R.Z., T.C. and R. Negrão prepared the visualizations. A.R.Z., T.C., W.L.E., F. Forest and W.J.B. wrote the original manuscript with support from A.A., S. Bellot, D.M.C., O.M.G., H. Sauquet and S.A.S. K.L., E.F., J.J.C., J.T.K., L. Pokorny, N.J.W., A.-Q.H., E.M.J., I.L., J.V., M.S.A., J.B.B., M.D.B., R.L.B., A.M.C.B., L. Broadhurst, A.B., D.J.C., M.M.C., M. W. Chase, L.W.C., M.J.M.C., P.-A.C., T.L.P.C., U.E., R.H.J.E., M. Estrella, M.F.F., P.L.F., M.G., L.J.G., S.W.G., J.H., T. Haevermans, J.C.H., O.H., T.R.H., K.K., E.A.K., M.A.K., F.L., C.J.L., D.-Z.L., S.L.-S., T.D.M., V.S.F.T.M., F.A.M., A.K.M., M.J.M., D.J.M., F.J.N., L. Palazzesi, J.P., D.S.P., O.A.P.-E., Y.P., G.M.P., R.K.R., R.R., H. Schaefer, A.S.-L., M.P.S., A.R.G.S., N.S., D.E.S., P.S.S., R. J. Soreng, P.F.S., S.C.K.S., A.H.T., T.M.A.U., J.M.V., J.J.W., T.C.W., Y.Y., S.Z. and I.J.L. reviewed the final manuscript. S.A.S., W.L.E., F. Forest and W.J.B. undertook supervision. P.J.K., I.J.L., F. Forest and W.J.B. acquired funding. V.M.A.B., P.J.K., I.J.L., F. Forest and W.J.B. were responsible for project administration.
Corresponding author
Correspondence to William J. Baker .
Ethics declarations
Competing interests.
The authors declare no competing interests.
Peer review
Peer review information.
Nature thanks Craig Barrett, Elena Kramer, Patrick Wincker and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended data fig. 1 tanglegram at ordinal level between this work (left) and the apg iv schematic tree (right)..
Branches colours represent the clades according to the composition proposed in each work. Posterior probability is presented only for nodes without maximum support. Coloured circles in the left tree represent the posterior probability of each node as: maximum (absent), between 1 and 0.95 (green), between 0.95 and 0.75 (yellow), between 0.75 and 0.5 (red), below 0.5 (black).
Extended Data Fig. 2 Comparison of node age estimates in the eight time-calibrated phylogenetic trees.
Each point represents a node and corresponds to the percentage difference in age estimates for that node between the two trees that are compared in each plot.
Extended Data Fig. 3 Comparison of stem ages of families and orders inferred in this study and Ramírez-Barahona et al. 5 .
a and b , Stem age comparison between our young tree (maximum constraint at the root node of 154 Ma) and the dataset CC_complete of Ramírez-Barahona et al. 5 . a , Ages in each study, coloured according to taxonomic rank and b , Age differences, calculated as age in this study minus age in Ramírez-Barahona et al. 5 c and d , Stem ages comparison between our old tree (maximum constraint at the root node of 247 Ma) and the dataset UC_complete from Ramírez-Barahona et al. 5 c , Ages in each study, coloured according to taxonomic rank and d , Age differences, calculated as age in this study minus age in Ramírez-Barahona et al. 5 .
Extended Data Fig. 4 Correlation between branch time duration and percentage of gene trees that do not share a congruent bipartition for the branch.
The results are based on the young tree (maximum constraint at the root node of 154 Ma). For each branch in the young tree, the percentage of gene trees that do not share a congruent bipartition with the species tree branch is plotted against the logarithm of the time duration for the branch.
Extended Data Fig. 5 Angiosperm-wide diversification and gene tree conflict through time.
This is equivalent to Fig. 3 but for the old tree (maximum constraint at the root node of 247 Ma). a , Estimated net diversification rate through time (yellow, left y-axis) and the level of gene tree conflict through time (blue, right y-axis). Net diversification rates are estimated with a model that enables speciation rates to vary between time intervals; the line is the posterior mean and the yellow shaded area is the 95% highest posterior density. Gene tree conflict is calculated from the percentage of gene trees that do not share a congruent bipartition with each species tree branch, with the plotted value being the mean across all species tree branches that cross each 2.5 Myr time slice. b , Cumulative percentage of extant orders and families that have originated through time. In both a and b, the background grey-scale gradient is the estimated percentage of extant lineages represented in the species tree through time (“sampling fraction”).
Extended Data Fig. 6 Summary of lineage-specific diversification rate shifts estimated by BAMM.
This is equivalent to Fig. 4 , but for the old tree (maximum constraint at the root node of 247 Ma). a , Diversification rate increases per lineage through time. The colour corresponds to the average magnitude of the rate increases during the time period. b , Equivalent to a, but for rate decreases. c , Equivalent to a, but focusing on the largest 25% of diversification rate increases. In a , b and c , the number of shifts is extracted from the maximum a posteriori shift configuration, the prior for the number of shifts is set to 10 and the background grey-scale gradient is the estimated percentage of extant lineages represented in the species tree through time (“sampling fraction”).
Supplementary information
Supplementary information.
Supplementary Methods, Results, Discussion, references, Figs. 1–24, captions for Tables 1–8, Files 1 and 2 and extended acknowledgements.
Reporting Summary
Peer review file, supplementary material, supplementary tables.
Supplementary Tables 1–8.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Zuntini, A.R., Carruthers, T., Maurin, O. et al. Phylogenomics and the rise of the angiosperms. Nature (2024). https://doi.org/10.1038/s41586-024-07324-0
Download citation
Received : 15 September 2023
Accepted : 15 March 2024
Published : 24 April 2024
DOI : https://doi.org/10.1038/s41586-024-07324-0
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.

Message placeholder

Recent Research Progress on Sustainable Energy Management System Based on Energy Efficiency and Renewable Energy
Erkata Yandri 1 ,2 * , Kukuh Priyo Pramono 1 , Very Sihombing 1 , Luqmanul Hakim Effendi 1 , Denis Ardianto 1 , Roy Hendroko Setyobudi 1 , Suherman Suherman 3 , Satriyo Krido Wahono 4 , Haryo Wibowo 5 , Marchel Putra Garfansa 6 and Afrida Rizka Farzana 7
1 Graduate School of Renewable Energy, Darma Persada University, Special Region of Jakarta 13450 2 Center of Renewable Energy Studies, Darma Persada University, Jakarta 13450, Indonesia 3 Diponegoro University, Semarang 50275, Central Java, Indonesia 4 Research Center for Food Technology and Processing (PRTPP), National Research and Innovation Agency (BRIN), Special Region of Yogyakarta 55861, Indonesia 5 Institute of Energy and Power Engineering, Zhejiang University of Technology, Hangzhou 310027, P.R. China 6 Universitas Islam Madura, Pamekasan 69317, Madura, East Java, Indonesia 7 IPB University, Bogor 16680, West Java, Indonesia
* Corresponding author: [email protected]
Energy Management Systems (EMS) have become increasingly important in efforts to address global energy challenges, such as increasing energy demand and climate change. EMS can be used to improve energy efficiency; reduce greenhouse gas emissions; and increase energy security. The purpose of the research is to review the latest research progress which focuses on EMS from various sectors based on energy efficiency and renewable energy. This research method involves four steps: selecting the EMS topic, searching for related papers using keywords on Google Scholar; summarizing and categorizing the obtained papers, and creating a table for easy understanding of the collected research; followed by analysis and discussion. As a result, recent research progress on sustainable EMS has been discussed, emphasizing categories like IoT; cloud data; controllers; reinforcement learning; renewable energy sources; energy storage; energy trading; and dashboards. The focus in EMS studies lies on IoT devices; controllers; reinforcement learning; and renewable energy; with less emphasis on energy trading and dashboards. The primary objective is to facilitate energy use tracking for users in various sectors, enabling them to assess efficiency and cost-effectiveness. This review facilitates energy tracking across diverse sectors for users, enabling evaluation of efficiency and cost-effectiveness.
Key words: Green energy system / home energy storage / IoT / smart grid / smart home
© The Authors, published by EDP Sciences, 2024

Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.

IMAGES
VIDEO
COMMENTS
Open access policy: Open Access Research Journal of Life Sciences (OARJLS) is an open access journal which means that all content is freely available without charge to the user or his/her institution. Users are allowed to read, download, copy, distribute, print, search, or link to the full texts of the articles in this journal without asking ...
Open Access Research Journal of Life Sciences (OARJLS) makes research outputs easy to publish, free to access, more visible through indexing. We're a not-for-profit organization that exists to make scholarly communications better. Open Access Research Journal of Life Sciences (OARJLS) do not take Publication fee for open access.
Life is an international, peer-reviewed, open access journal of scientific studies related to fundamental themes in life sciences, from basic to applied research, published monthly online by MDPI.The Astrobiology Society of Britain (ASB) and Spanish Association for Cancer Research (ASEICA) are affiliated with Life and their members receive a discount on the article processing charges.
Open Life Sciences is an open-access, peer-reviewed journal committed to encompassing a wide array of subjects within the life sciences. It is poised to captivate a diverse readership across the fields of biology, biotechnology, and medicine, including scholars who would like to gain profound insights into the latest scientific breakthroughs, as well as non-scholarly readers, who seek to ...
Aims. Life (ISSN 2075-1729) is an international, peer-reviewed scientific open access journal concerned with fundamental themes in life sciences, from basic to applied research. Our aim is to encourage scientists to publish their experimental and theoretical results in as much detail as possible. There is no restrictions on the maximum length ...
Life Sciences cater to the research and discovery of living organisms, and include areas of study such as Agricultural and Biological Sciences, Immunology and Neuroscience. Publish Open Access Open access lies at the core of Elsevier's publishing mission - in fact, today, almost all of our journals offer open access options.
Open access information. Life Sciences offers authors two choices to publish their research: Gold open access. Subscription. Articles are freely available to both subscribers and the wider public with permitted reuse. Articles are made available to subscribers as well as developing countries and patient groups through our access programs.
It is a Referred, Indexed, Online International Journal. Open Access Research Journal of Life Sciences (OARJLS) is published as a Quarterly Journal with 4 issues per year. Open Access Research Journal of Life Sciences (OARJLS) offers fast publication of quality Research and Review articles.
About the directory. DOAJ is a unique and extensive index of diverse open access journals from around the world, driven by a growing community, and is committed to ensuring quality content is freely available online for everyone. DOAJ is committed to keeping its services free of charge, including being indexed, and its data freely available.
All Life is the home for publishing multidisciplinary reproducible life science research, led by a team of subject experts. As an international open access journal, we aim to break down siloed specialisms, bringing together all experts to foster better collaboration and information sharing across the life sciences. As part of our Elevate Series authors will receive a concierge-level publishing ...
Biology and Life Sciences Forum. Biology and Life Sciences Forum is an open access journal dedicated to publishing findings resulting from conferences, workshops, and similar events, in all areas of biology, life sciences and at the interface of related disciplines. The conference organizers and proceedings editors are responsible for managing ...
Open Access in Medicine and Life Sciences. In medicine and the life sciences, open access is particularly supported by mandates from research funders such as the National Institutes of Health (NIH), the Wellcome Trust, and the Bill & Melinda Gates Foundation. Calls for free access to the results of research in medicine and the life sciences ...
London, UK.SAGE, one of the world's leading independent and academic publishers, has today announced that it has launched the Journal of International Life Sciences Research(JILSR), a new open access (OA) journal dedicated to the rapid publication of high-quality articles in the form of life sciences research and review papers.. As a companion to the Journal of International Medical Research ...
The Harvard Open Journals Program will offer publishing and hosting services to help the Harvard community launch new open access journals, or to convert existing journals to open access. The program will offer two support models: an overlay model which takes advantage of open access repositories, such as Harvard's DASH , and a brand-new ...
Life Sciences is an international journal publishing articles that emphasize the molecular, cellular, and functional basis of therapy. The journal emphasizes the understanding of mechanism that is relevant to all aspects of human disease and translation to patients. All articles are rigorously …. View full aims & scope.
Research Journal of Life Science is an open access publishes papers three times a year on April, August and December. The main objective of Research Journal of Life Science is to provide a platform for the international scholars, academicians and researchers to share the contemporary thoughts and innovation in the fields of life science. Research Journal of Life Science aims to promote studies ...
In its 2019-2023 Strategic Plan, the Institut Pasteur confirms its long-term strategy to "promote open access to publications and research data."In 2021, in line with this commitment, it published a Charter for Open Access to Scientific Publications. In the charter, the Institut Pasteur calls for all its scientific publications (research articles, reviews, letters, books, conference papers and ...
This discussion between Joseph Lerro, Open Research Business Development Manager with Taylor & Francis, and Rachel Scott, Associate Dean for Information Assets at Illinois State University (ISU) and Editor of Library Resources & Technical Services , explores one such partnership that led to an innovative, social-sciences focused OA agreement.
Journal of Biophotonics, ... Ideal curricula should bridge scientific and engineering tools with expertise in photonics and life sciences to perform biomedical research on molecular, cellular, and tissue level using bio-inspired photonic devices ... Open access funding provided by IReL.
About the Journal. Research & Reviews : A Journal of Life Sciences (rrjols): 2249-8656 (e) is a peer-reviewed hybrid open-access journal launched in 2011 focused on the rapid publication of View Full Focus and Scope….
Journal of Innovative Research in Life Sciences (JIRLS) is an open access, peer reviewed online and print journal, aimed at publishing latest research findings in life sciences.The Journal comprise techniques suitable in promoting the dissemination of research findings that is expected to be of benefit to the basic needs in health, agriculture, biotechnology, pharmaceutical and food industries.
Open Access Research Journal of Life Sciences (OARJLS) publishes manuscripts (Original research, review articles, Short communication and letter to editor) on original work, either experimental or theoretical) from all aspects of Life Sciences (Biology, Genetics, Biological Anthropology, Botany, Medical Sciences, Veterinary Sciences, Biochemical Genetics, Biometry, Clinical Genetics ...
Phylogenomic analysis of 7,923 angiosperm species using a standardized set of 353 nuclear genes produced an angiosperm tree of life dated with 200 fossil calibrations, providing key insights ...
Based on the 2018 Life Experience Survey of Indonesian Children and Adolescents, ... Australian Journal of Forensic Sciences Volume 56, 2024 - Issue sup1: Extended Abstracts of the ... Open access. Overview; Open journals; Open Select; Dove Medical Press; F1000Research; Opportunities.
Heidi Mertes g Department of Philosophy and Moral Sciences and Department of Public Health and Primary Care, Ghent University ... This study was funded by the Practice Oriented Scientific Research (PWO) of VIVES University of Applied Sciences. ... Open access. Overview; Open journals; Open Select; Dove Medical Press; F1000Research ...
Open Access Research Journal of Life Sciences ISSN 2783-025X (Online) Navigation. Home; About us. About the Journal; Editorial Board; Indexing; Journal policies; Publication ethics; Editorial Board; View Articles. Issue in progress; Current Issue; Archive Past Issues; Guide for Author. Instructions for Authors;
Life, an international, peer-reviewed Open Access journal. Next Volume Volume 14 (2024) Previous Volume Volume 12 (2022) ... The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal. Original Submission Date ... Life, Volume 13 (2023) Vol. 13, Iss. 1 January 2023. Table of Contents.
This research method involves four steps: selecting the EMS topic, searching for related papers using keywords on Google Scholar; summarizing and categorizing the obtained papers, and creating a table for easy understanding of the collected research; followed by analysis and discussion.